 Back
BackChapter 6
Study Guide - Smart Notes

Hemoglobin Function and Oxygen Transport
Oxygen Binding and Release by Hemoglobin
Hemoglobin is a tetrameric protein responsible for transporting oxygen from the lungs to tissues and facilitating its release based on tissue needs. The oxygen dissociation curve illustrates the relationship between the partial pressure of oxygen and hemoglobin saturation, reflecting hemoglobin's cooperative binding properties.
Oxygen Release: In a resting body, hemoglobin typically releases about 25% of its carried oxygen, meaning on average, one oxygen molecule is released per hemoglobin tetramer, leaving it about 75% saturated.
Cooperativity: Hemoglobin exhibits cooperative binding, where the binding of one oxygen molecule increases the affinity for subsequent oxygen molecules.
Regulation: Factors such as pH (Bohr effect), 2,3-Bisphosphoglycerate (2,3-BPG), and CO2 levels modulate hemoglobin's oxygen affinity, allowing adaptation to varying physiological conditions.
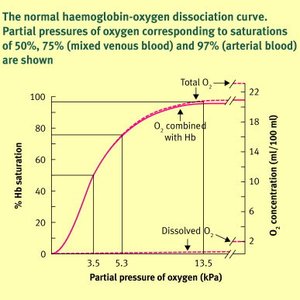
Additional info: The sigmoidal shape of the curve is a hallmark of cooperative binding, distinguishing hemoglobin from myoglobin, which has a hyperbolic curve due to its non-cooperative binding.
Chapter 6: Enzymes
What Are Enzymes?
Enzymes are biological catalysts, primarily globular proteins, that accelerate chemical reactions without being consumed. Some RNA molecules (ribozymes) also possess catalytic activity.
Specificity: Enzymes exhibit high specificity for their substrates, minimizing side reactions.
Reaction Conditions: Enzymes function under mild physiological conditions (pH ~7, 37°C).
Regulation: Enzyme activity can be regulated, allowing control over metabolic pathways.
Enzyme-Substrate Complex and Selectivity
Enzymes bind substrates at their active sites, forming an enzyme-substrate (ES) complex. This interaction is crucial for catalytic specificity and efficiency.
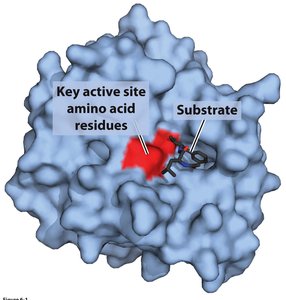
Active Site: The region where substrate binding and catalysis occur, often involving key amino acid residues.
Substrate Selectivity: Only specific substrates fit into the active site, ensuring precise catalysis.
Enzyme Catalysis and Reaction Coordinate
Enzymes accelerate reactions by lowering the activation energy (ΔG‡) required to reach the transition state, without altering the overall equilibrium (ΔG').
Activation Energy (ΔG‡): The energy barrier that must be overcome for a reaction to proceed.
Transition State: A high-energy, unstable state during the conversion of substrate to product.
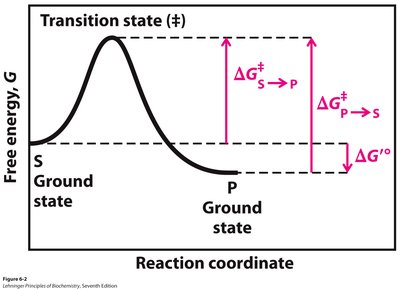
Additional info: The difference in free energy between the ground state and the transition state determines the reaction rate.
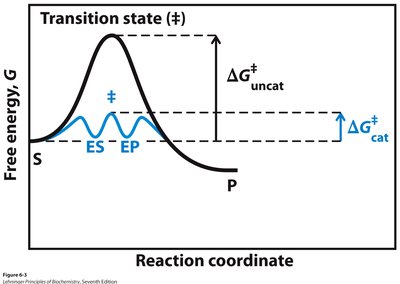
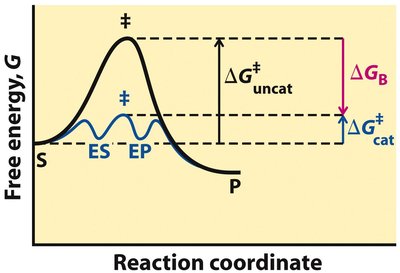
Enzyme-Catalyzed vs. Uncatalyzed Reactions
Enzyme-catalyzed reactions proceed through lower activation energies compared to uncatalyzed reactions, often involving transient intermediates (ES, EP).
Mechanisms of Enzyme Catalysis
Enzymes utilize several catalytic mechanisms to enhance reaction rates:
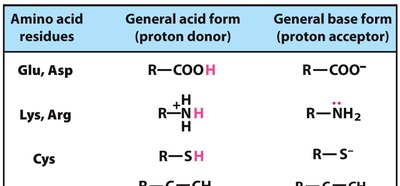
Acid-Base Catalysis: Transfer of protons to stabilize intermediates.
Covalent Catalysis: Formation of transient covalent bonds with substrates.
Metal Ion Catalysis: Use of metal ions to facilitate redox reactions or substrate orientation.
Electrostatic Catalysis: Stabilization of charged transition states.
Binding Energy and Transition State Stabilization
Enzymes achieve rate enhancement by stabilizing the transition state more than the substrate, utilizing binding energy (ΔGB).
Proximity and Orientation: Enzymes bring reactive groups into close proximity and proper orientation, reducing entropy and facilitating reaction.
Transition State Binding: Enzyme active sites are complementary to the transition state, not the substrate, ensuring efficient catalysis.
Enzyme Kinetics
Basic Kinetic Model
The simplest enzyme-catalyzed reaction can be represented as:
First-Order Reaction: Rate depends on substrate concentration:
Second-Order Reaction: For two substrates:
Michaelis-Menten Kinetics
The Michaelis-Menten equation describes the relationship between substrate concentration and reaction velocity for many enzymes:
Vmax: Maximum velocity when the enzyme is saturated with substrate.
Km: Substrate concentration at which the reaction velocity is half-maximal; reflects enzyme affinity for substrate.
kcat: Turnover number, the number of substrate molecules converted to product per enzyme per second.
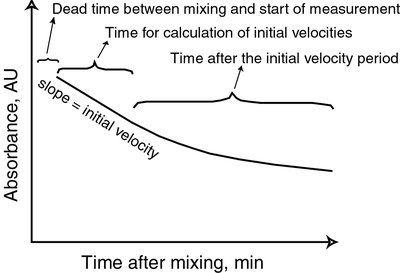
Additional info: At low [S], velocity is proportional to [S]; at high [S], velocity approaches Vmax and becomes independent of [S].
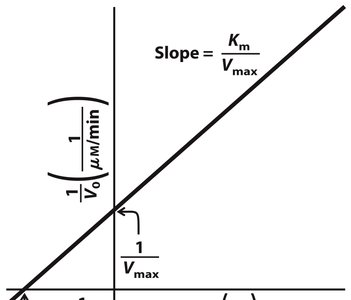
Lineweaver-Burk Plot
The Lineweaver-Burk (double-reciprocal) plot linearizes the Michaelis-Menten equation for easier determination of kinetic parameters:
Slope:
Y-intercept:
X-intercept:
Enzyme Inhibition
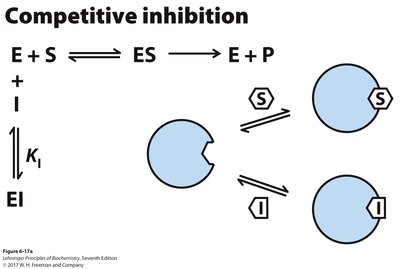
Enzyme inhibitors decrease enzyme activity and are classified as reversible or irreversible. Reversible inhibitors include competitive, uncompetitive, and mixed inhibitors, each affecting kinetic parameters differently.
Competitive Inhibition: Inhibitor binds to the active site, increasing apparent Km but not affecting Vmax.
Uncompetitive Inhibition: Inhibitor binds only to the ES complex, decreasing both Km and Vmax.
Mixed Inhibition: Inhibitor binds to both enzyme and ES complex, affecting both Km and Vmax.
Two-Substrate Reactions
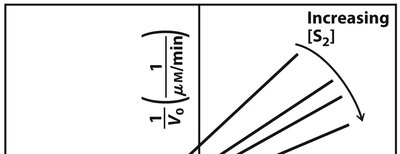
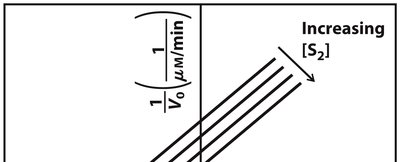
Enzyme-catalyzed reactions involving two substrates can follow sequential (ordered or random) or ping-pong (double-displacement) mechanisms. Lineweaver-Burk plots help distinguish these mechanisms based on the intersection or parallelism of lines.
Sequential Mechanism: Both substrates must bind before any product is released; lines intersect on the plot.
Ping-Pong Mechanism: One product is released before the second substrate binds; lines are parallel on the plot.
Summary Table: Types of Enzyme Inhibition
Type | Binding Site | Effect on Vmax | Effect on Km | Lineweaver-Burk Plot |
|---|---|---|---|---|
Competitive | Active site | No change | Increases | Lines intersect at y-axis |
Uncompetitive | ES complex | Decreases | Decreases | Lines are parallel |
Mixed | Enzyme or ES complex | Decreases | Varies | Lines intersect left of y-axis |
