 Back
BackDNA Replication, Repair, and Mutagenesis: Biochemistry Study Notes
Study Guide - Smart Notes

DNA Replication: Fundamental Concepts
Modes of DNA Replication
DNA replication is the process by which a cell duplicates its DNA, ensuring genetic information is passed to daughter cells. Three theoretical models were proposed for DNA replication: conservative, semi-conservative, and dispersive. Experimental evidence supports the semi-conservative model, where each daughter DNA molecule consists of one parental and one newly synthesized strand.
Conservative replication: The parental DNA remains intact, and a completely new copy is made.
Semi-conservative replication: Each daughter DNA contains one old (parental) and one new strand.
Dispersive replication: Parental and new DNA are interspersed in both strands.
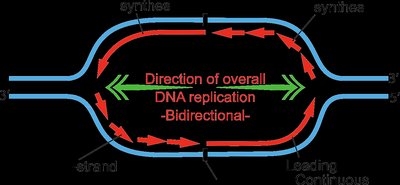
Bidirectional and Semi-Discontinuous Replication
DNA replication proceeds in both directions from the origin, forming two replication forks. The leading strand is synthesized continuously, while the lagging strand is synthesized discontinuously in short fragments (Okazaki fragments).
Leading strand: Synthesized continuously in the 5' to 3' direction.
Lagging strand: Synthesized discontinuously as Okazaki fragments, later joined by DNA ligase.
Enzymes and Proteins in DNA Replication
Key Enzymes in E. coli DNA Replication
Multiple enzymes and proteins coordinate the accurate and efficient replication of DNA:
DNA Polymerase III: The main replicative polymerase in E. coli, composed of multiple subunits (α: polymerase activity, ε: 3'→5' exonuclease proofreading, θ: unknown function).
DNA Gyrase (Topoisomerase): Introduces negative supercoils to relieve torsional strain.
Helicase: Unwinds the DNA double helix using ATP.
Primase: Synthesizes short RNA primers required for DNA polymerase to initiate synthesis.
DNA Ligase: Seals nicks between Okazaki fragments.
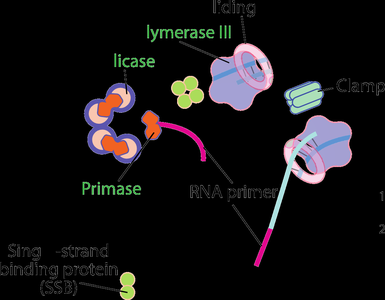

Initiation and Termination of Replication
Replication begins at specific origins and ends at defined termination sites:
Initiation: dnaA protein binds to the origin, causing local DNA unwinding and assembly of the replication machinery.
Termination: In E. coli, the Tus protein binds to terminator sequences, halting the replication fork.
Comparison of DNA Replication in E. coli and Humans
Replication mechanisms are conserved, but there are key differences between prokaryotes and eukaryotes:
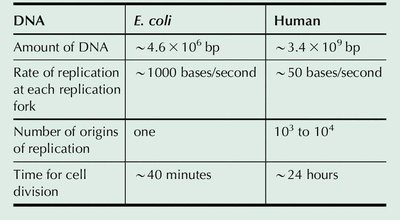
DNA | E. coli | Human |
|---|---|---|
Amount of DNA | ~4.6 × 106 bp | ~3.4 × 109 bp |
Rate of replication at each fork | ~1000 bases/second | ~50 bases/second |
Number of origins of replication | one | 103 to 104 |
Time for cell division | ~40 minutes | ~24 hours |
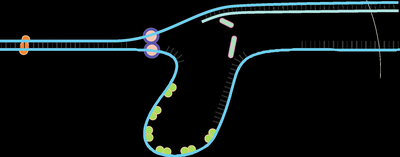
Mechanisms of Leading and Lagging Strand Synthesis
Elongation Process
DNA synthesis occurs in the 5' to 3' direction. The leading strand is synthesized continuously, while the lagging strand is synthesized in Okazaki fragments, which are later joined.
Okazaki fragments: Short DNA fragments synthesized on the lagging strand.
Primer removal: DNA polymerase I removes RNA primers and fills the gaps with DNA.
Nick sealing: DNA ligase joins the fragments to form a continuous strand.

Specialized Replication Mechanisms
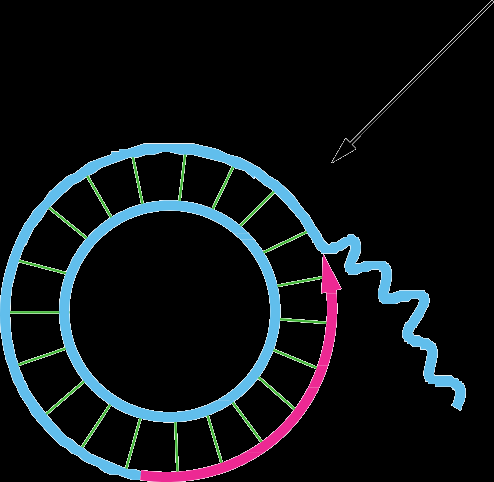
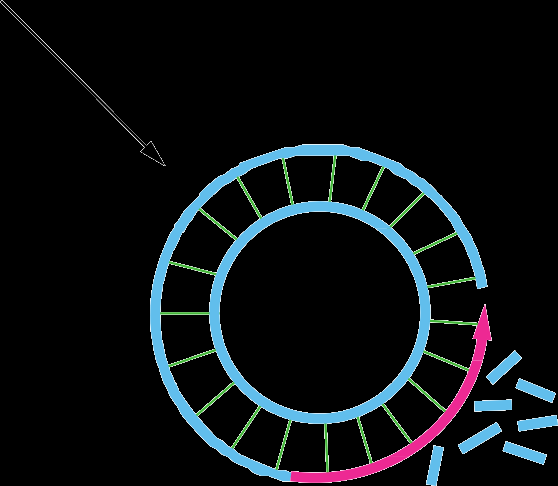
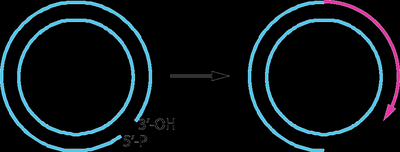
Rolling-Circle Replication
This mechanism is used by some viruses and during bacterial conjugation. It involves continuous synthesis of a new strand while the template strand is displaced.
Phage DNA and bacterial mating: Rolling-circle replication allows rapid and repeated copying of circular DNA.
Primer not required: The leading strand remains attached to the template.

Replication of Eukaryotic Chromosomes
Regulation and Complexity
Eukaryotic DNA replication is tightly regulated and involves multiple origins of replication, nucleosome disassembly/reassembly, and telomere maintenance.
Multiple origins: Each chromosome has many origins to ensure timely replication.
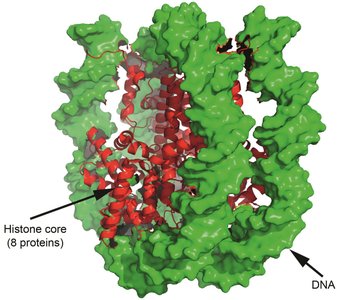
Histone complexes: DNA is packaged into nucleosomes, requiring coordinated histone synthesis and assembly during replication.
Telomeres: Specialized structures at chromosome ends, maintained by telomerase.
Inhibitors of DNA Replication
Classes and Mechanisms
Several inhibitors are used in research and therapy to target DNA replication:
Precursor synthesis inhibitors: Block nucleotide biosynthesis.
Template/priming inhibitors: Interfere with DNA or RNA primer formation.
Polymerase inhibitors: Directly inhibit DNA polymerases (e.g., acyclovir, aphidicolin).
DNA-damaging agents: Cause strand breaks or cross-links (e.g., bleomycin, mitomycin).
DNA Mutations and Repair
Types of Mutations
Mutations are changes in the DNA sequence and can be classified by size and effect:
Small-scale mutations: Base substitutions (transitions, transversions), insertions, deletions.
Large-scale mutations: Chromosomal rearrangements (translocations, inversions, deletions, nondisjunction).
Consequences: Silent, missense, and nonsense mutations affect protein coding differently.
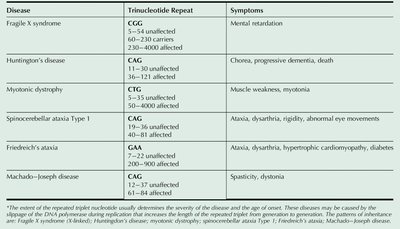
Inherited Diseases Involving Trinucleotide Repeats
Expansion of trinucleotide repeats in certain genes leads to inherited disorders.
Disease | Trinucleotide Repeat | Symptoms |
|---|---|---|
Fragile X syndrome | CGG | Mental retardation |
Huntington's disease | CAG | Chorea, progressive dementia, death |
Myotonic dystrophy | CTG | Muscle weakness, myotonia |
Spinocerebellar ataxia Type 1 | CAG | Ataxia, dysarthria, rigidity, abnormal eye movements |
Friedreich's ataxia | GAA | Ataxia, dysarthria, hypertrophic cardiomyopathy, diabetes |
Machado–Joseph disease | CAG | Spasticity, dystonia |
Ames Test for Mutagenicity
The Ames test is a widely used assay to assess the mutagenic potential of chemical substances. It uses a strain of Salmonella typhimurium that cannot synthesize histidine. If a chemical induces a back mutation, the bacteria survive in histidine-deficient media, indicating mutagenic (and potentially carcinogenic) properties.
Types of DNA Damage
Base loss: Loss of a base from the DNA backbone.
Base modification: Deamination, alkylation, oxidation.
Replication errors: Mismatches, insertions, deletions.
Inter-strand crosslinks: Covalent links between DNA strands.
DNA-protein crosslinks: Covalent links between DNA and proteins.
Strand breaks: Single- or double-strand breaks from radiation or chemicals.
DNA Repair Systems
Categories of DNA Repair
Cells employ several mechanisms to maintain genetic stability:
Direct reversal: Enzymatic reversal of specific types of damage (e.g., photolyase for thymine dimers, MGMT for O6-methylguanine).
Excision repair: Removal of damaged DNA followed by resynthesis (mismatch, base excision, nucleotide excision repair).
Damage tolerance: Allows replication to proceed past unrepaired lesions (e.g., SOS response in bacteria).



Human Diseases Associated with DNA Repair Deficiency
Xeroderma Pigmentosum (XP): Defective nucleotide excision repair, leading to extreme UV sensitivity and cancer risk.
Ataxia telangiectasia: Defective response to ionizing radiation, neurological symptoms.
Hereditary nonpolyposis colon cancer (HNPCC): Defective mismatch repair, increased cancer risk.
Fanconi’s anemia: Defective repair of interstrand cross-links, bone marrow failure.
Werner syndrome: Accelerated aging due to defective DNA helicase.
Bloom’s syndrome: Defective BLM helicase, increased chromosomal instability.
Cockayne’s syndrome: Defective transcription-coupled repair, growth and neurological defects.
Glioblastoma: Dysfunction in direct reversal repair (MGMT).
