 Back
BackEnzyme Catalysis and Carbohydrate Structure: Biochemistry Study Guide
Study Guide - Smart Notes

Enzyme Catalysis
General Principles of Catalysis
Enzymes are biological catalysts that accelerate chemical reactions by lowering the activation energy required for the reaction to proceed. They achieve this by stabilizing the transition state and providing an alternative reaction pathway.
Catalyst: A substance that increases the rate of a chemical reaction without being consumed or permanently changed.
Activation Energy (Ea): The minimum energy required for a reaction to occur.
Transition State Stabilization: Enzymes bind and stabilize the transition state, reducing the energy barrier.
Equilibrium: Enzymes do not affect the position of equilibrium; they only increase the rate at which equilibrium is reached.
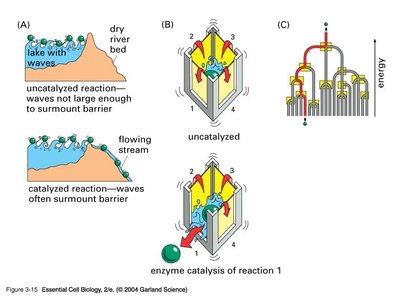
Mechanisms of Enzyme Catalysis
Enzyme catalysis involves both entropic and enthalpic factors. The enzyme positions and orients substrates to facilitate the formation of the transition state, and may distort the substrate or active site to further reduce activation energy.
Entropic Effects: Enzymes bring substrates into close proximity and proper orientation, reducing the entropy cost of reaction.
Enthalpic Effects: Strong binding interactions between enzyme and substrate in the transition state provide favorable enthalpy.
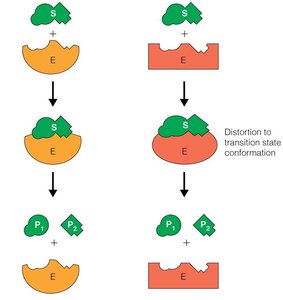
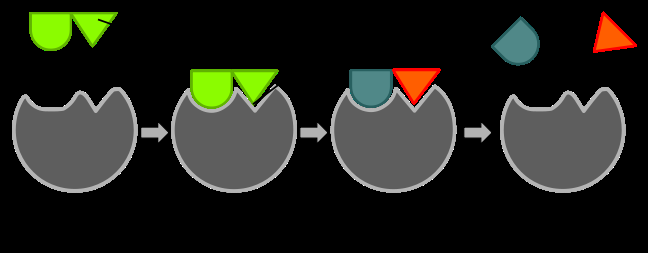
Models of Enzyme-Substrate Interaction
Two main models describe how enzymes bind substrates:
Lock-and-Key Model: The enzyme's active site is complementary to the substrate's shape.
Induced-Fit Model: The enzyme changes shape upon substrate binding, increasing affinity for the transition state and stabilizing it.
Michaelis-Menten Kinetics
The Michaelis-Menten equation describes the rate of enzyme-catalyzed reactions:
Michaelis-Menten Equation:
Vmax: Maximum reaction rate at saturating substrate concentration.
Km: Substrate concentration at which the reaction rate is half-maximal; reflects enzyme affinity for substrate.
Kcat: Turnover number; maximum number of substrate molecules converted to product per enzyme per second.
Amino Acids in Enzyme Active Sites
Specific amino acids in enzyme active sites play critical roles in catalysis, often with altered pKa values due to their environment.
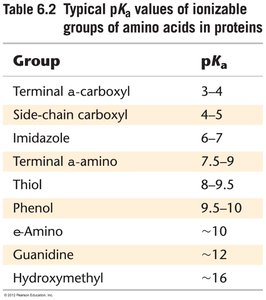
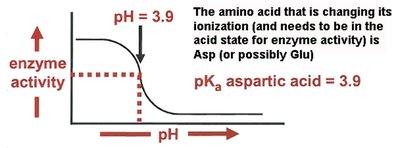
Effect of pH on Enzyme Activity
Enzyme activity is sensitive to pH, which affects the ionization state of amino acid side chains in the active site. The pKa of these groups determines their protonation state and thus their catalytic function.
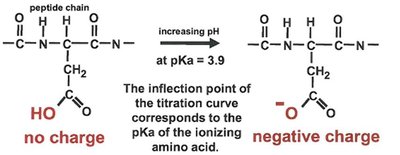
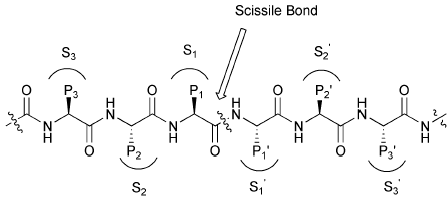
Serine Protease Mechanism
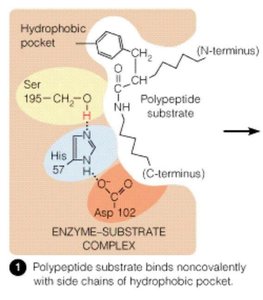
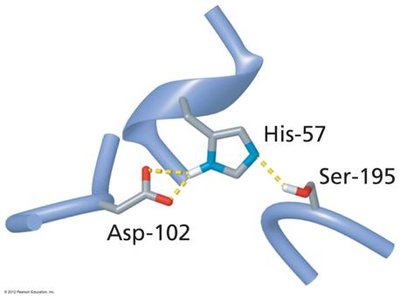
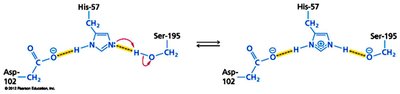
Serine proteases, such as chymotrypsin, utilize a catalytic triad (Ser, His, Asp) to cleave peptide bonds. The substrate binds in the active site, and the triad facilitates nucleophilic attack and stabilization of the transition state.
Enzyme Inhibition
Types of Reversible Inhibition
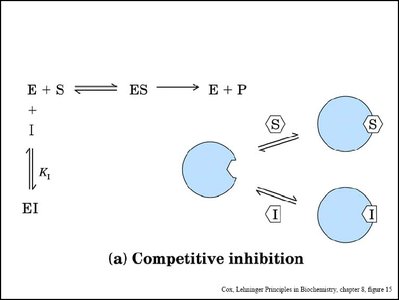
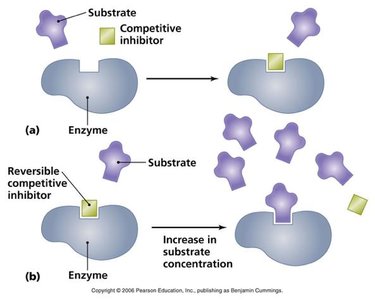
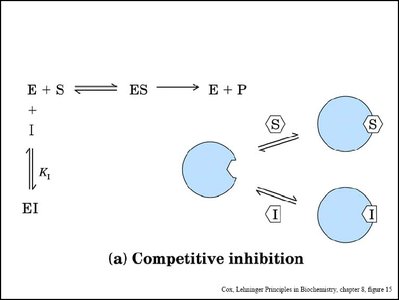
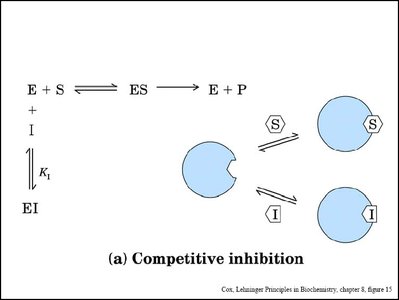
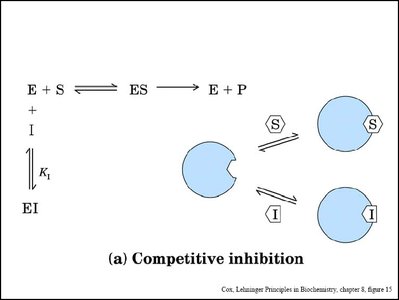
Enzyme inhibitors reduce enzyme activity by binding to the enzyme or enzyme-substrate complex. Three main types are competitive, noncompetitive (mixed), and uncompetitive inhibition.
Competitive Inhibition: Inhibitor binds to the active site, preventing substrate binding. Increases apparent Km, Vmax unchanged.
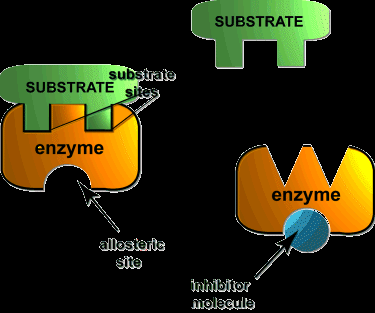
Noncompetitive (Mixed) Inhibition: Inhibitor binds to enzyme or ES complex. Decreases Vmax, Km may change.
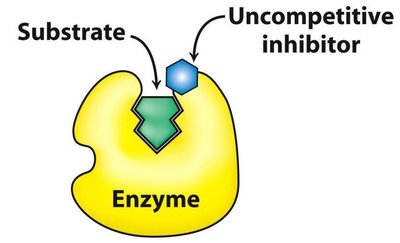
Uncompetitive Inhibition: Inhibitor binds only to ES complex. Both Km and Vmax decrease.
Regulation of Enzyme Activity
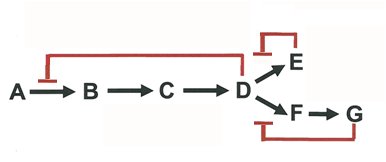
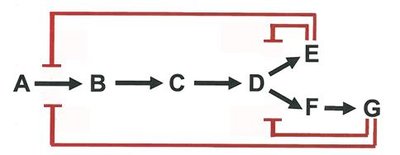
Feedback and End-Product Inhibition
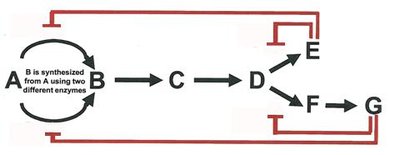
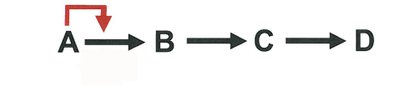
Metabolic pathways are regulated by feedback inhibition, where the end product inhibits an early enzyme in the pathway, conserving resources and energy.
Coenzymes and Essential Ions
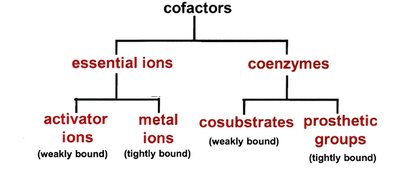
Cofactors in Enzyme Catalysis
Many enzymes require cofactors for activity. Cofactors include essential ions and coenzymes, which may be tightly or loosely bound.
Metal-Activated Enzymes
Metal ions such as Mg2+ are required for the activity of many enzymes, including kinases, by stabilizing substrate binding and facilitating catalysis.
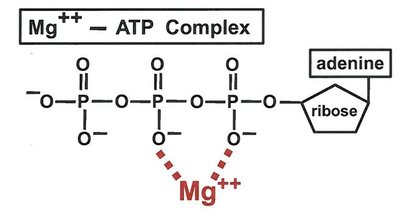
Carbohydrate Structure
Classification of Sugars
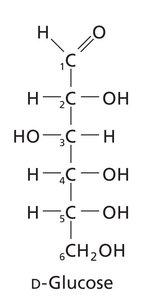
Carbohydrates are classified based on their functional groups (aldoses vs ketoses), number of carbons, and chirality. Monosaccharides can be aldoses (aldehyde group) or ketoses (ketone group).
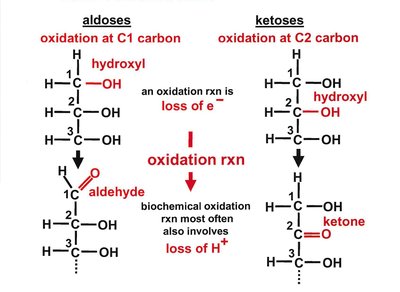
Chirality and Stereochemistry
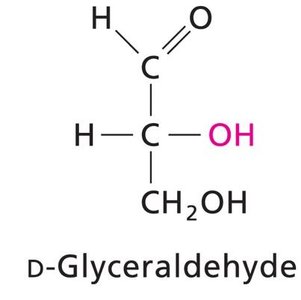
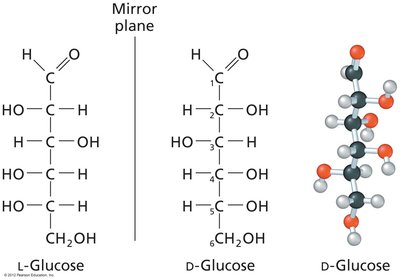
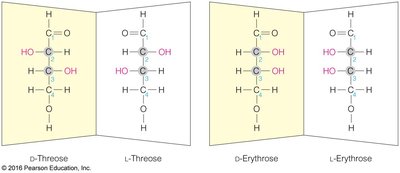
Monosaccharides exhibit chirality, with D and L forms determined by the configuration of the chiral carbon farthest from the carbonyl group. Enantiomers are mirror images, while diastereomers differ at one or more chiral centers.
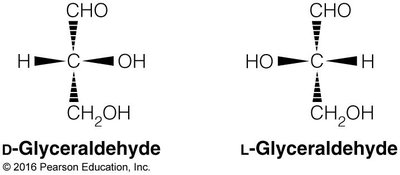
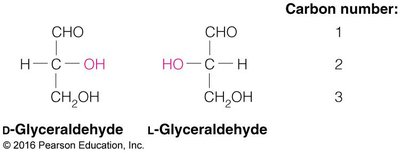
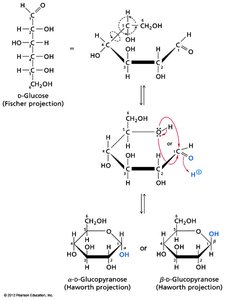
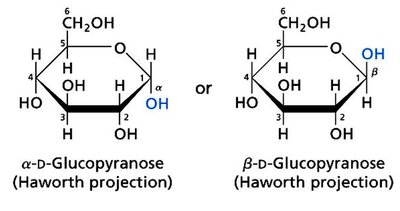
Cyclization and Anomeric Carbon
Monosaccharides can cyclize to form rings, generating a new chiral center at the anomeric carbon. The orientation of the OH group at the anomeric carbon determines the α or β form.
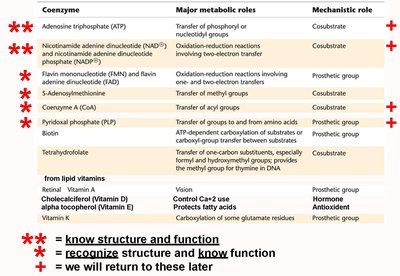
Summary Table: Major Coenzymes and Their Functions
Coenzyme | Major Metabolic Role | Mechanistic Role |
|---|---|---|
ATP | Transfer of phosphoryl or nucleotidyl groups | Cosubstrate |
NAD(P) | Oxidation-reduction reactions | Cosubstrate |
FMN/FAD | Oxidation-reduction reactions | Prosthetic group |
Coenzyme A | Transfer of acyl groups | Cosubstrate |
Biotin | ATP-dependent carboxylation | Prosthetic group |
Tetrahydrofolate | Transfer of one-carbon groups | Prosthetic group |
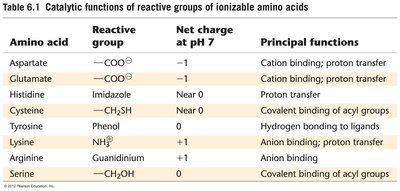
Summary Table: Catalytic Functions of Ionizable Amino Acids
Amino Acid | Reactive Group | Net Charge at pH 7 | Principal Functions |
|---|---|---|---|
Aspartate | COO- | -1 | Cation binding; proton transfer |
Glutamate | COO- | -1 | Cation binding; proton transfer |
Histidine | Imidazole | Near 0 | Proton transfer |
Cysteine | CH2SH | Near 0 | Covalent binding of acyl groups |
Tyrosine | Phenol | 0 | Hydrogen bonding to ligands |
Lysine | NH3+ | +1 | Anion binding; proton transfer |
Arginine | Guanidinium | +1 | Anion binding |
Serine | CH2OH | 0 | Covalent binding of acyl groups |
Additional info:
Expanded explanations of enzyme catalysis, inhibition, and carbohydrate stereochemistry were added for completeness.
Tables were recreated from the original materials for clarity and study purposes.
