 Back
BackEnzyme Catalysis, Mechanisms, and Carbohydrate Structure: Mini-Textbook Study Notes
Study Guide - Smart Notes

Enzyme Catalysis and Mechanisms
General Principles of Catalysis
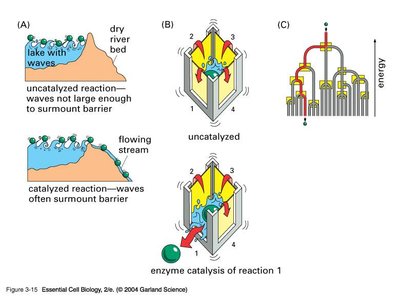
Catalysts are substances that increase the rate of a chemical reaction without being consumed or permanently altered. Enzymes, biological catalysts, are essential for accelerating biochemical reactions in cells, which would otherwise proceed too slowly to sustain life.
Activation Energy Reduction: Catalysts lower the activation energy (Ea), increasing the fraction of molecules able to reach the transition state.
Equilibrium Position: Catalysts do not affect the position of equilibrium; they only speed up the attainment of equilibrium.
Transition State Stabilization: Enzymes stabilize the transition state, facilitating the conversion of substrate (S) to product (P).
Enzyme Catalysis Mechanisms
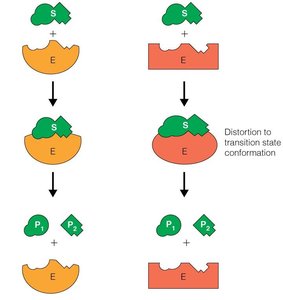
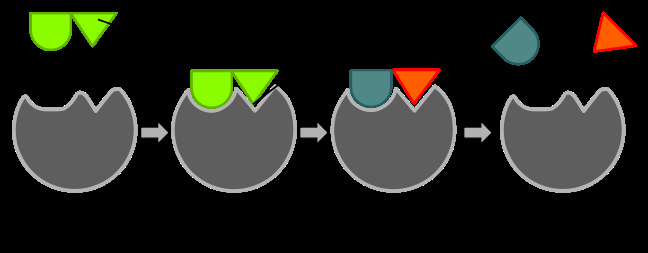
Enzyme catalysis involves binding substrates, orienting them, and stabilizing transition states. The induced fit model is favored, where initial weak binding induces conformational changes in the enzyme, strengthening substrate binding and transition state stabilization.
Entropic Factors: Enzymes position and orient substrates, reducing entropy and increasing reaction rates.
Enthalpic Factors: Strong binding interactions in the transition state provide favorable enthalpy.
Induced Fit Model: Substrate binding induces enzyme conformational changes, increasing affinity for the transition state.
Michaelis-Menten Kinetics
The Michaelis-Menten equation describes the rate of enzyme-catalyzed reactions as a function of substrate concentration:
Equation:
Parameters: Km is the substrate concentration at half-maximal velocity; Vmax is the maximum rate at saturating substrate.
Kcat: Turnover number, maximum number of substrate molecules converted per enzyme per second.
Catalytic Efficiency: , measures enzyme specificity and efficiency.
Amino Acids in Enzyme Active Sites
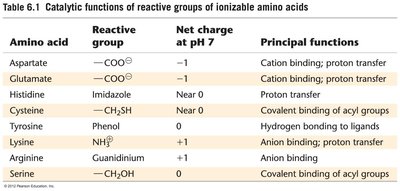
Amino acids in enzyme active sites often have altered pKa values due to their microenvironment, affecting their catalytic roles. Common active site residues include Asp, Glu, His, Lys, Cys, Tyr, and Ser.
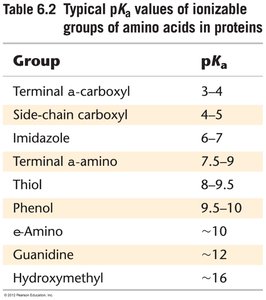
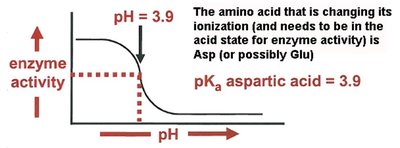
Effect of pH on Enzyme Activity
Enzyme activity is highly dependent on pH, which affects the ionization state of active site residues. Titration curves can reveal the involvement of specific amino acids in catalysis.
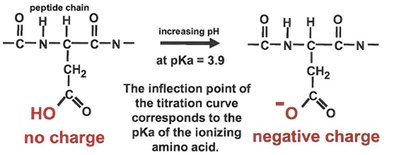
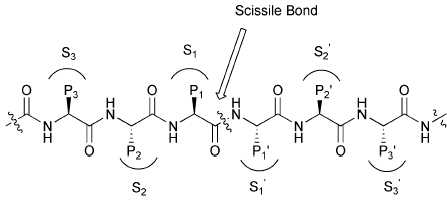
Enzyme Reaction Mechanisms
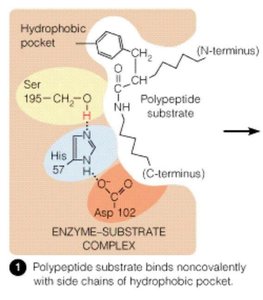
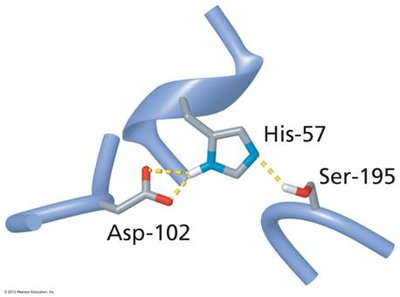
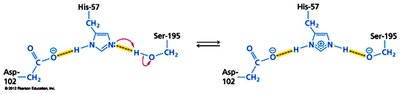
Enzyme-catalyzed reactions often involve nucleophilic/electrophilic attacks, acid-base catalysis, and covalent intermediates. The serine protease mechanism is a classic example, involving a catalytic triad (Ser, His, Asp).
Nucleophilic/Electrophilic Reactions: Functional groups donate or accept electrons.
Acid-Base Catalysis: Amino acid side chains act as proton donors/acceptors.
Covalent Catalysis: Formation of covalent enzyme-substrate intermediates.
Enzyme Inhibition
Types of Reversible Inhibition
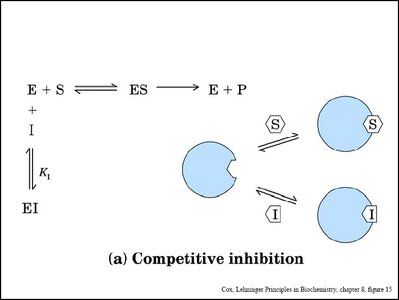
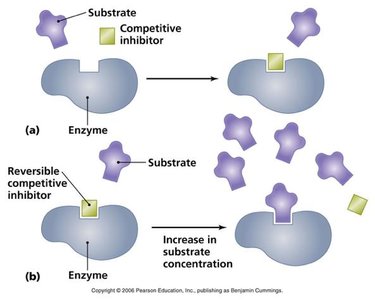
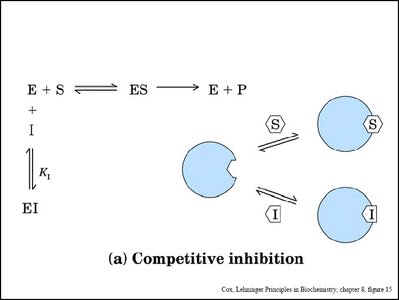
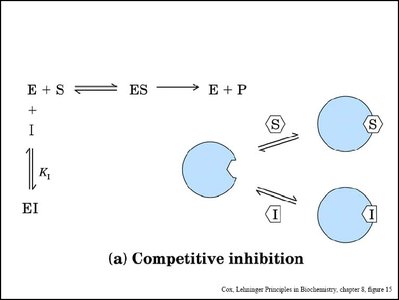
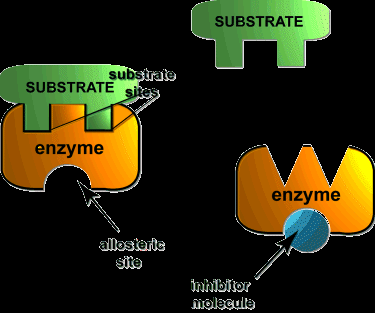
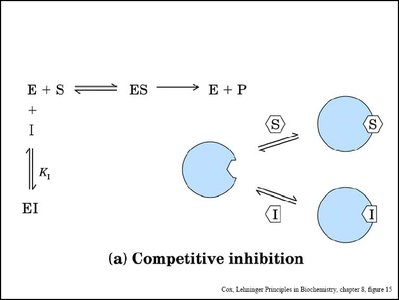
Enzyme inhibitors can decrease enzyme activity by binding to the enzyme or enzyme-substrate complex. The main types are competitive, noncompetitive (mixed), and uncompetitive inhibition.
Competitive Inhibition: Inhibitor binds to the active site, preventing substrate binding. Increases apparent Km, Vmax unchanged.
Noncompetitive (Mixed) Inhibition: Inhibitor binds to enzyme or ES complex, decreases Vmax, Km may change.
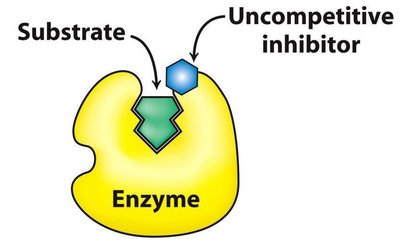
Uncompetitive Inhibition: Inhibitor binds only to ES complex, decreases both Km and Vmax by same factor.
Regulation of Enzyme Activity
Feedback and Complex Inhibition
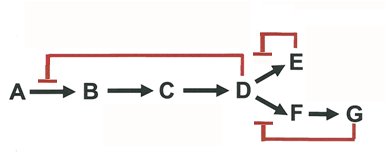
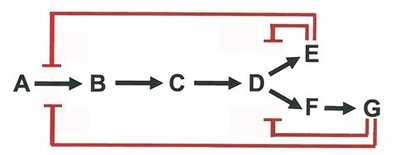
Enzyme activity is regulated by feedback inhibition and more complex patterns, ensuring efficient metabolic control.
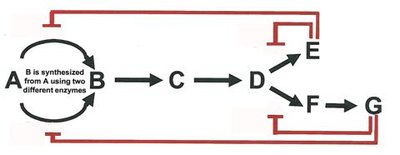
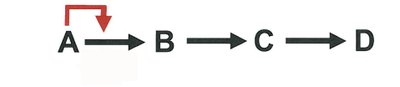
Feedback Inhibition: End-product inhibits the first enzyme in a pathway.
Sequential and Concerted Inhibition: Multiple products regulate different steps.
Differential Inhibition: Multiple enzymes are regulated by different effectors.
Coenzymes and Essential Ions
Cofactor Classification and Functions
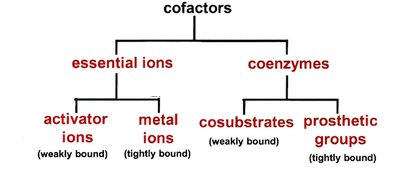
Many enzymes require cofactors for activity, which can be essential ions or coenzymes. Cofactors are classified by their binding and function.
Essential Ions: Activator ions (weakly bound), metal ions (tightly bound).
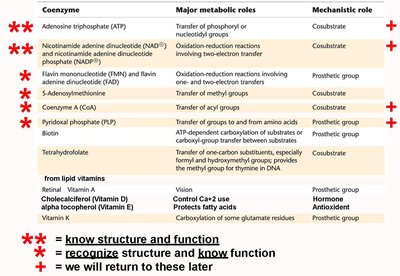
Coenzymes: Cosubstrates (weakly bound), prosthetic groups (tightly bound).
Metal-Activated Enzymes
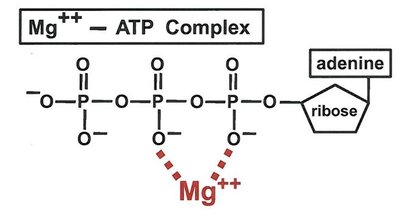
Metal ions such as Mg2+ are required for substrate binding and catalysis in many enzymes, e.g., kinases.
Carbohydrate Structure and Stereochemistry
Classification and Structure of Sugars
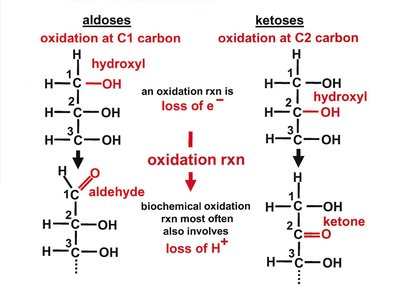
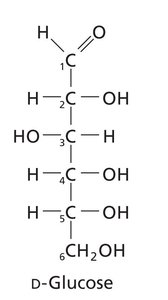
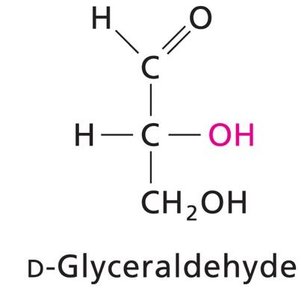
Carbohydrates are classified as monosaccharides, disaccharides, oligosaccharides, and polysaccharides. Monosaccharides are further divided into aldoses (aldehyde group) and ketoses (ketone group).
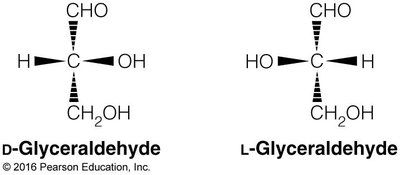
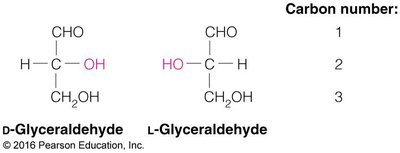
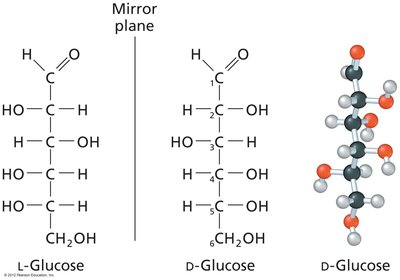
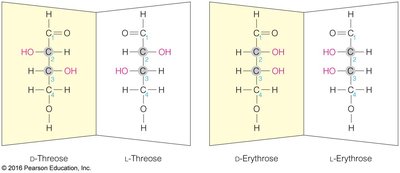
Chirality: Most carbohydrates are D-chiral; L-chiral forms are rare in nature.
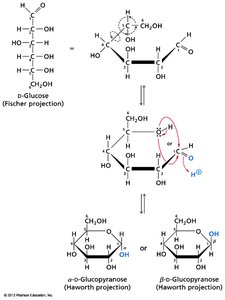
Fischer and Haworth Projections: Used to represent sugar structures and ring forms.
Cyclization and Anomeric Carbon
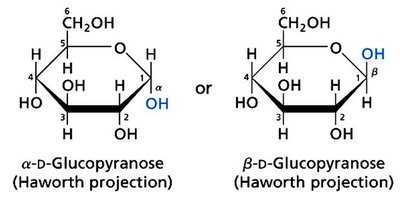
Monosaccharides cyclize to form rings, generating a new chiral center at the anomeric carbon. The orientation of the OH group at the anomeric carbon defines α and β forms.
Polysaccharides and Glycosidic Bonds
Polysaccharides are classified as homoglycans (one type of sugar) or heteroglycans (multiple types). Glycosidic bonds link monosaccharides, and their configuration (α or β) and position determine polysaccharide properties.
Storage Homoglycans: Starch (amylose, amylopectin), glycogen.
Structural Homoglycans: Cellulose, chitin.
Heteroglycans: Glycosaminoglycans, proteoglycans, glycoproteins, glycolipids.
Nucleic Acids: Structure and Function
Nucleotide Structure
Nucleotides are composed of a phosphate group, a pentose sugar (ribose or deoxyribose), and a nitrogenous base (purine or pyrimidine). Nucleic acids are polymers of nucleotides linked by phosphodiester bonds.
Purines: Adenine, guanine (two rings).
Pyrimidines: Cytosine, thymine, uracil (one ring).
Phosphodiester Bonds: Link 3' OH of one nucleotide to 5' phosphate of the next.
DNA and RNA Structure
DNA is typically double-stranded, forming a right-handed B-DNA helix. RNA is usually single-stranded but can form complex secondary and tertiary structures.
Antiparallel Strands: DNA strands run in opposite directions.
Base Pairing: Hydrogen bonds between complementary bases (A-T, G-C).
Chromatin Packaging: DNA is wrapped around histone proteins to form nucleosomes in eukaryotes.
RNA Types and Functions
RNA molecules serve informational, structural, and catalytic roles. Major types include mRNA, rRNA, tRNA, miRNA, and ribozymes.
mRNA: Messenger RNA, encodes proteins.
rRNA: Ribosomal RNA, forms ribosome core.
tRNA: Transfer RNA, delivers amino acids during translation.
miRNA/siRNA: Regulate gene expression.
Ribozymes: RNA molecules with catalytic activity.
UV Absorbance and DNA Quantification
Nucleic acids absorb UV light at 260 nm due to their conjugated double bonds. This property is used to quantify DNA and RNA concentration and purity.
Beer’s Law:
DNA: 1 A260 = 50 μg/ml
RNA: 1 A260 = 40 μg/ml
DNA Melting and Stability
DNA stability depends on base composition, solvent conditions, cation concentration, denaturants, and temperature. GC-rich DNA is more stable due to stronger base stacking.
