 Back
BackEnzyme Kinetics and Regulation: A Comprehensive Study Guide
Study Guide - Smart Notes

Biochemical Reactions and Enzyme Catalysis
Introduction to Biochemical Reactions
Biochemical reactions are chemical processes that occur within living cells, forming the basis of metabolism and life. These reactions are typically organized into metabolic pathways and are characterized by two essential features: they are enzyme-driven and exergonic (energy-releasing).
Enzyme-driven: All biochemical reactions are catalyzed by specific enzymes, which accelerate reaction rates by lowering activation energy.
Exergonic: Reactions proceed in one direction, ensuring energy flow and metabolic irreversibility.
Understanding biochemical reactions requires addressing two fundamental questions: How fast does the reaction occur? (kinetics) and In which direction does the reaction proceed? (thermodynamics).
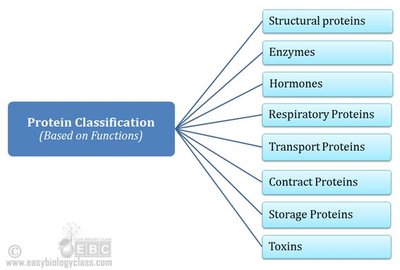
Classification of Proteins by Function
Proteins serve diverse roles in biological systems, including catalysis, structure, transport, and defense.
Structural proteins: e.g., keratin, collagen
Enzymes (catalytic proteins): e.g., hexokinase, pepsin
Hormonal proteins: e.g., insulin
Transport proteins: e.g., hemoglobin
Receptor proteins: e.g., hormone receptors
Contractile proteins: e.g., actin, myosin
Storage proteins: e.g., ovalbumin
Genetic proteins: e.g., nucleoproteins
Defense proteins: e.g., immunoglobulins
Enzyme Structure and Function
Definition and Properties of Enzymes
Enzymes are primarily proteins (with some RNA-based exceptions called ribozymes) that act as biological catalysts. They accelerate specific chemical reactions without being consumed or permanently altered in the process. Enzymes lower the activation energy required for reactions but do not affect the overall Gibbs free energy change.
Active site: The region on the enzyme where substrates bind and reactions occur.
Allosteric site: A regulatory site distinct from the active site, where modulators can bind and alter enzyme activity.
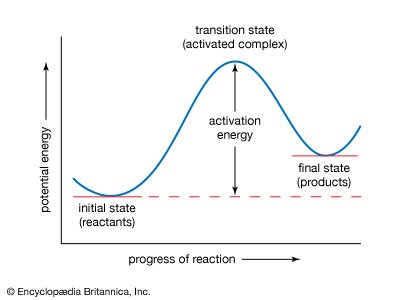
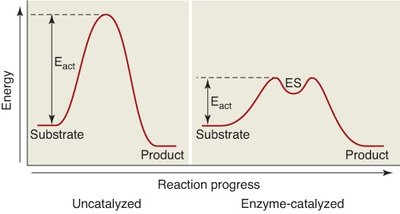
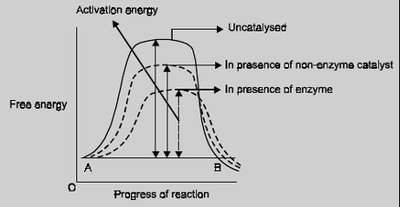
Enzyme Catalysis and Activation Energy
Enzymes facilitate reactions by stabilizing the transition state and reducing the activation energy barrier, thereby increasing the reaction rate.
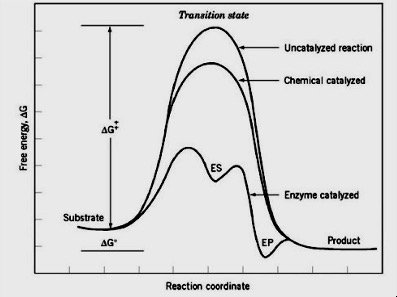
Enzyme Kinetics
Reaction Rate and Kinetics
The rate of an enzyme-catalyzed reaction depends on substrate concentration, enzyme concentration, and the kinetic parameters of the enzyme. The reaction rate is often described by the Michaelis-Menten equation for single-subunit enzymes.
Reaction rate (v): Change in product or substrate concentration per unit time.
Initial velocity (V0): The fastest rate, measured at the very start of the reaction.
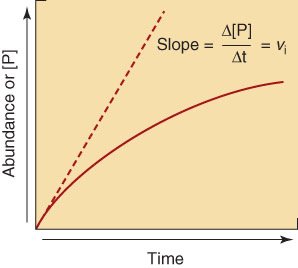
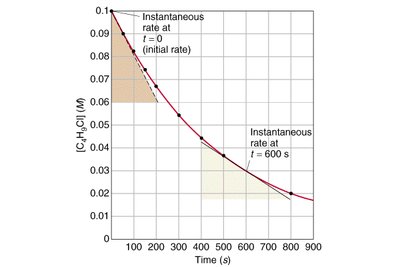
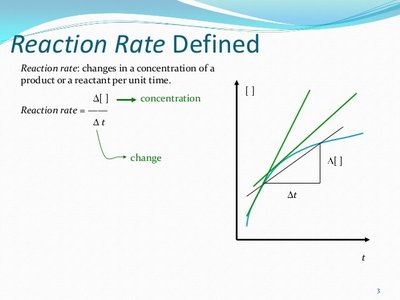
Arrhenius Equation and Reaction Rate Constant
The Arrhenius equation relates the reaction rate constant (k) to activation energy (EA) and temperature (T):
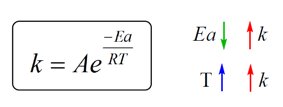
Michaelis-Menten Kinetics
The Michaelis-Menten equation describes the relationship between substrate concentration and reaction velocity for many enzymes:
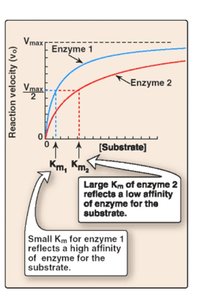
Vmax: Maximum reaction velocity at saturating substrate concentration.
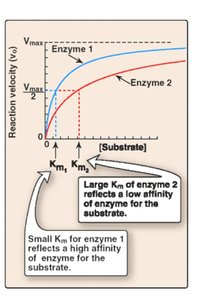
Km: Michaelis constant; substrate concentration at which the reaction rate is half of Vmax. Lower Km indicates higher substrate affinity.
Catalytic efficiency: Measured by the ratio Vmax/Km.
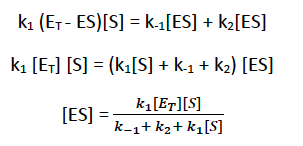
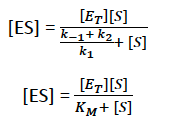
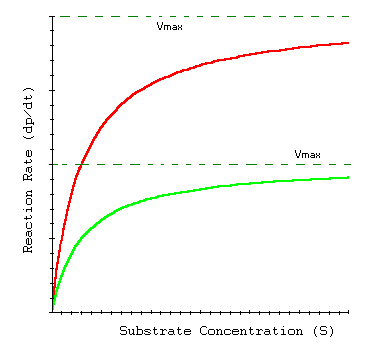
Interpretation of Kinetic Parameters
Km: Indicates binding affinity; lower values mean tighter binding.
Vmax: Indicates binding capacity; higher values mean more active sites or more enzyme present.
Vmax/Km: Indicates catalytic efficiency; higher values mean more efficient enzymes.
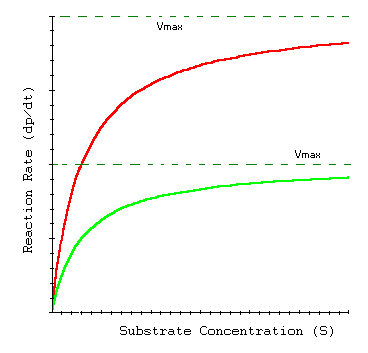
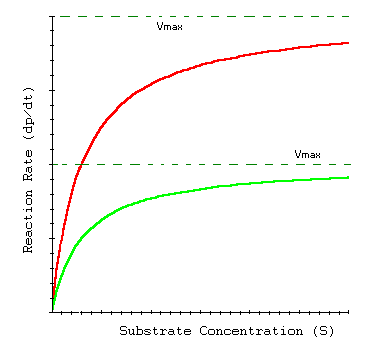
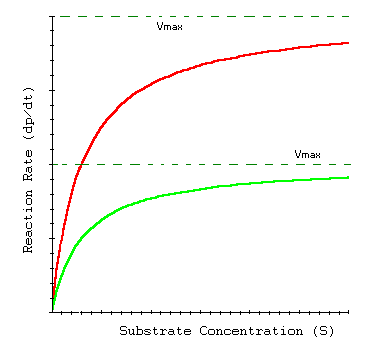
Enzyme Kinetics in Transporters
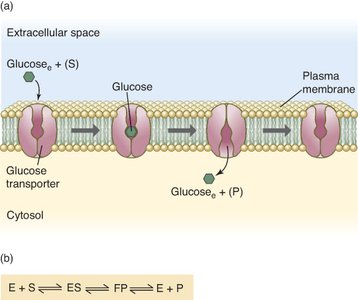
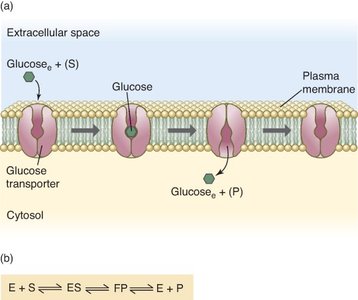
Membrane transport proteins, such as glucose transporters, often follow Michaelis-Menten kinetics, reflecting their enzyme-like substrate binding and transport properties.
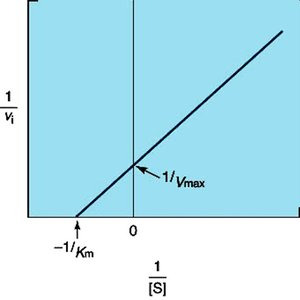
Lineweaver-Burk (Double-Reciprocal) Plot
The Lineweaver-Burk plot linearizes the Michaelis-Menten equation by plotting 1/v versus 1/[S], allowing easier determination of Km and Vmax from the intercepts.

Enzyme Regulation and Inhibition
Regulation of Enzyme Activity
Enzyme activity can be regulated by altering enzyme concentration, modifying enzyme structure, or through the action of activators and inhibitors.
Up-regulation: Increases [enzyme]total and Vmax.
Down-regulation: Decreases [enzyme]total and Vmax.
Allosteric regulation: Modulators bind to allosteric sites, changing enzyme conformation and activity.
Types of Enzyme Inhibition
Enzyme inhibitors are classified based on their interaction with the enzyme:
Reversible inhibition: Inhibitor binds non-covalently and can dissociate.
Irreversible inhibition: Inhibitor binds covalently, permanently inactivating the enzyme.
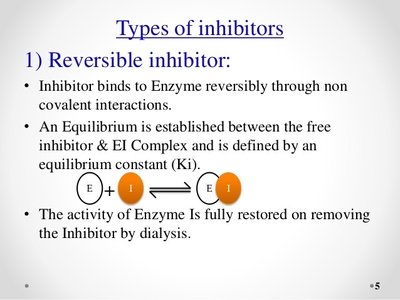

Reversible Inhibition: Competitive, Uncompetitive, and Noncompetitive
Inhibition Type | Effect on Vmax | Effect on Km | Effect on Vmax/Km |
|---|---|---|---|
Competitive | No change | Increases | Decreases |
Uncompetitive (Anticompetitive) | Decreases | Decreases | No change |
Noncompetitive (Mixed) | Decreases | No change | Decreases |
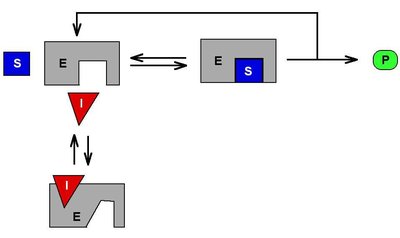
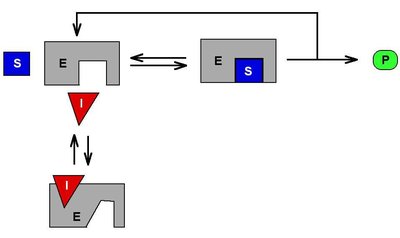
Competitive inhibition: Inhibitor competes with substrate for the active site. Vmax is unchanged, Km increases.
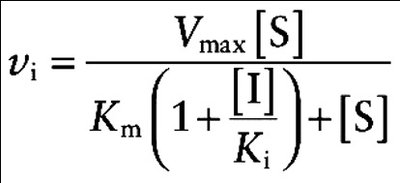
Allosteric Enzymes and Hill Kinetics
Allosteric Regulation and Cooperativity
Allosteric enzymes have multiple subunits and active sites. They exhibit cooperative binding, where substrate binding at one site affects binding at others. This is described by the Hill equation:
n > 1: Positive cooperativity
n = 1: No cooperativity (reduces to Michaelis-Menten)
n < 1: Negative cooperativity
Enzyme Classification
Seven Major Classes of Enzymes (EC System)
EC Number | Class | Reaction Catalyzed | Example |
|---|---|---|---|
EC1 | Oxidoreductase | Oxidation-reduction | Lactate dehydrogenase |
EC2 | Transferase | Group transfer | Hexokinase |
EC3 | Hydrolase | Hydrolysis | Sucrase |
EC4 | Lyase | Addition/removal of groups | Aldolase |
EC5 | Isomerase | Isomerization | Glucose phosphate isomerase |
EC6 | Ligase | Bond formation with ATP hydrolysis | Pyruvate carboxylase |
EC7 | Translocase | Transport across membranes | ADP/ATP translocase |
Summary
Enzymes are essential biological catalysts, classified by their function and reaction type.
Enzyme kinetics, described by Michaelis-Menten and Hill equations, provides insight into enzyme efficiency and regulation.
Enzyme activity is regulated by substrate concentration, enzyme concentration, and inhibitors (competitive, uncompetitive, noncompetitive, irreversible).
Understanding enzyme kinetics and regulation is fundamental for biochemistry, pharmacology, and biotechnology.
