 Back
BackEnzymes and Catalysis: Structure, Function, and Regulation
Study Guide - Smart Notes

Enzymes and Catalysis
Introduction to Enzymes
Enzymes are biological catalysts that accelerate biochemical reactions without being consumed in the process. They are primarily proteins, though some RNA molecules (ribozymes) also exhibit catalytic activity. Enzymes are essential for life, enabling reactions to occur at rates compatible with cellular function.
Catalyst: A substance that increases the rate of a chemical reaction without undergoing permanent change.
Enzyme: A protein (or RNA) that acts as a catalyst in biological systems.
Dynamic Nature: Enzymes are not static; their structure and function can be influenced by environmental factors and regulatory molecules.
Protein Structure and Function
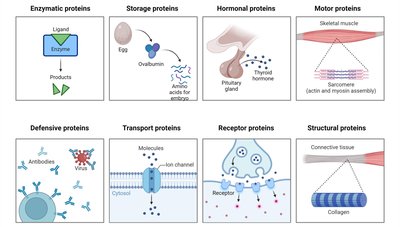
Proteins can be classified as fibrous or globular, each with distinct structural and functional properties. Enzymes are typically globular proteins, which allows for complex folding and formation of active sites.
Fibrous Proteins: Extended, repetitive structure; generally insoluble; function in support and protection.
Globular Proteins: Compact, irregular structure; generally soluble; function in catalysis, transport, signaling, and immunity.
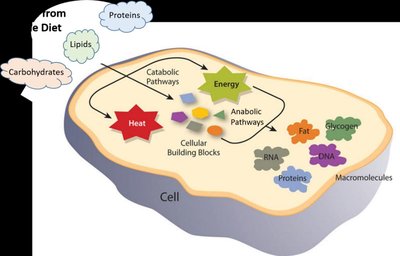
Metabolism and Enzymes
Metabolism encompasses all chemical reactions in a cell, divided into catabolic (breakdown) and anabolic (synthesis) pathways. Enzymes facilitate these reactions, ensuring efficient energy transfer and macromolecule synthesis.
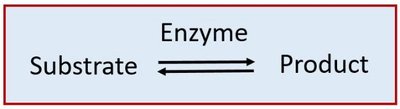
Stages and Components of Enzymatic Reactions
An enzymatic reaction involves the binding of substrate(s) to the enzyme's active site, conversion to product(s), and release. The process is reversible and governed by thermodynamic principles.
Substrate: The molecule upon which an enzyme acts.
Product: The molecule(s) formed from the substrate.
Active Site: The region of the enzyme where substrate binding and catalysis occur.
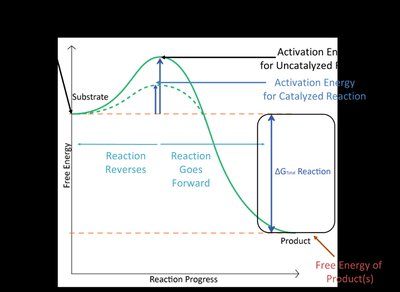
Thermodynamics of Enzyme Catalysis
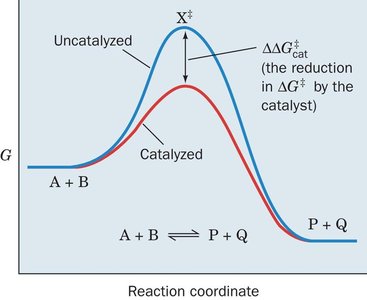
Enzymes lower the activation energy (ΔG‡) required for a reaction, increasing the reaction rate. The overall free energy change (ΔG) of the reaction remains unchanged by catalysis.
ΔG: Change in free energy; determines reaction spontaneity.
ΔG‡: Activation energy; determines reaction rate.
Transition State (X‡): The highest energy state during the reaction.
Enzyme Efficiency and Specificity
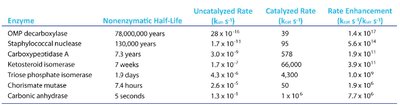
Enzymes are highly efficient, often increasing reaction rates by factors of 106 to 1012 compared to uncatalyzed reactions. They operate under mild conditions and exhibit high specificity for their substrates.
Reaction Rate Enhancement: Orders of magnitude higher than chemical catalysts.
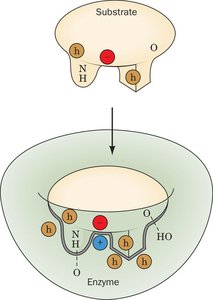
Specificity: Enzymes recognize substrates based on shape (geometric complementarity) and charge (electronic complementarity).
Enzyme-Substrate Interaction Models
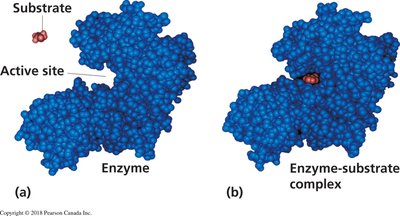
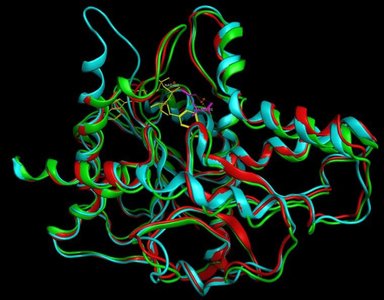
Two primary models describe how enzymes interact with substrates: the lock-and-key model and the induced fit model. The lock-and-key model suggests a rigid active site, while the induced fit model proposes that the enzyme adapts its shape upon substrate binding.
Lock-and-Key Model: Substrate fits into the active site without structural change.
Induced Fit Model: Enzyme changes shape to accommodate the substrate.

Enzyme Nomenclature and Classification
The International Union of Biochemistry and Molecular Biology (IUBMB) classifies enzymes based on the reactions they catalyze. Each enzyme is assigned an Enzyme Commission (EC) number, reflecting its class and specific activity.
Six Major Classes: Oxidoreductases, Transferases, Hydrolases, Lyases, Isomerases, Ligases.
EC Number: Hierarchical identifier for enzyme classification.
Accepted Name: Commonly used name.
Systematic Name: Describes substrate(s) and reaction type.
EC Class | Reaction Type |
|---|---|
1. Oxidoreductases | Oxidation-reduction |
2. Transferases | Group transfer |
3. Hydrolases | Hydrolysis |
4. Lyases | Bond cleavage (non-hydrolytic) |
5. Isomerases | Isomerization |
6. Ligases | Bond formation (with ATP hydrolysis) |

Bioinformatic Resources for Enzyme Study
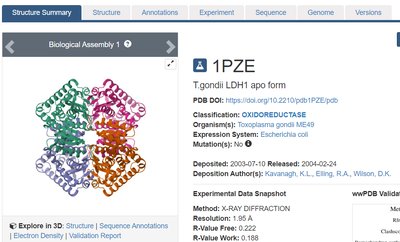
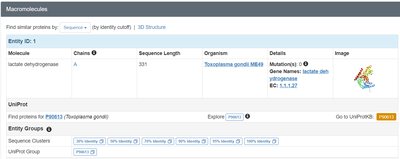
Databases such as BRENDA and the Protein Data Bank (PDB) provide detailed information on enzyme structure, function, and classification. These resources are essential for research and understanding enzyme mechanisms.

Cofactors and Enzyme Function
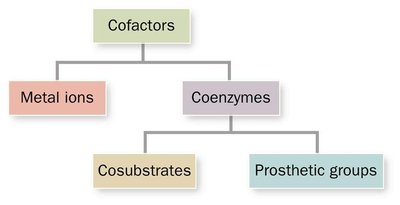
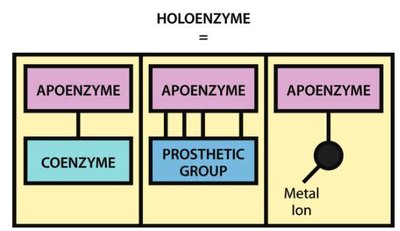
Many enzymes require non-protein molecules called cofactors for activity. Cofactors can be metal ions or organic molecules (coenzymes), which may be tightly or loosely bound to the enzyme.
Metal Ions: Essential for catalytic activity in some enzymes.
Coenzymes: Organic molecules that assist in enzyme function; can be cosubstrates or prosthetic groups.
Holoenzyme: The active enzyme with its cofactor.
Apoenzyme: The protein portion without the cofactor.
Environmental Effects on Enzyme Activity
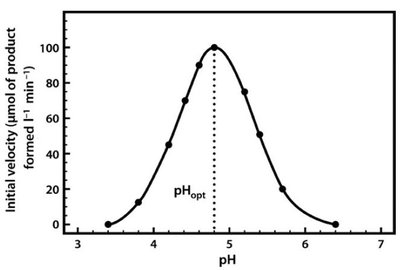
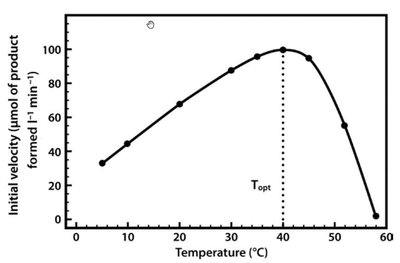
Enzyme activity is sensitive to temperature and pH. Each enzyme has an optimal temperature and pH at which it functions most efficiently. Deviations from these conditions can lead to denaturation and loss of activity.
Temperature: Increases reaction rate up to a point; excessive heat denatures the enzyme.
pH: Alters ionization of amino acids; extreme pH can denature the enzyme.

Enzyme Regulation and Inhibition
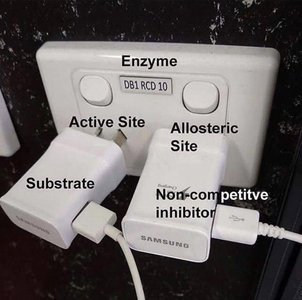
Enzymes are regulated by various mechanisms, including competitive, allosteric, and feedback inhibition. Regulation ensures proper metabolic control and prevents wasteful reactions.
Competitive Inhibition: Inhibitor binds to the active site, blocking substrate access.
Allosteric Inhibition: Inhibitor binds to a site other than the active site, altering enzyme conformation.
Feedback Inhibition: End product of a pathway inhibits an earlier enzyme.
Summary of Learning Outcomes
Describe enzymatic reactions using biochemical terminology.
Classify and name enzymes according to IUBMB standards.
Explain enzyme-substrate interactions and models.
Use thermodynamic arguments to explain enzyme catalysis.
Interpret reaction coordinate diagrams.
Evaluate effects of regulators and environmental factors on enzyme activity.
Additional info:
Enzyme kinetics and inhibition are covered in greater detail in subsequent lessons.
Bioinformatic resources are increasingly important for modern biochemistry research.
