 Back
BackIntroduction to Proteins: Structure, Function, and Sequence
Study Guide - Smart Notes

Introduction to Proteins
Amino Acids: The Building Blocks of Proteins
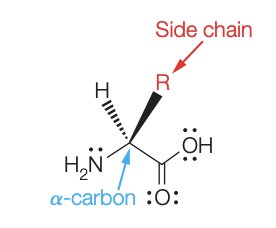
Amino acids are organic molecules that serve as the monomers for all proteins. Each amino acid contains a central α-carbon bonded to an amino group, a carboxyl group, a hydrogen atom, and a unique side chain (R group) that determines its properties.
α-amino acids are the standard monomers found in proteins.
The side chain (R group) varies among amino acids and is responsible for their chemical diversity.
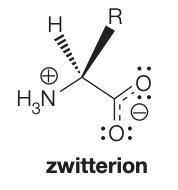
At physiological pH (~7), amino acids exist as zwitterions, carrying both positive and negative charges.
Classification of Amino Acids
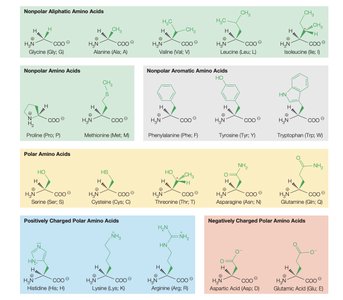
Amino acids are classified based on the properties of their side chains:
Nonpolar aliphatic
Nonpolar aromatic
Polar uncharged
Positively charged (basic)
Negatively charged (acidic)
Peptide Bonds and Protein Structure
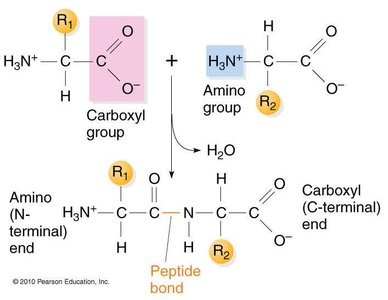
Proteins are formed by linking amino acids through peptide bonds (amide bonds). The peptide bond is formed between the carboxyl group of one amino acid and the amino group of another, releasing water in a condensation reaction.
Peptide bonds exhibit planar geometry due to partial double-bond character, restricting rotation.
Peptide bonds can exist in trans or cis configurations, with trans being more common due to reduced steric hindrance.
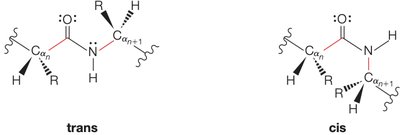
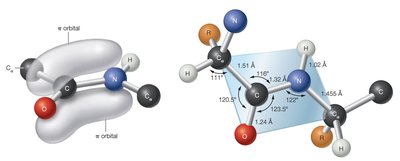
Formation and Stability of Peptide Bonds
The formation of peptide bonds is thermodynamically unfavored in aqueous solution, with a free energy change of approximately +10 kJ/mol. Polypeptides are metastable and hydrolyze rapidly only under extreme conditions or in the presence of catalysts such as proteases or strong acids (e.g., 6 M HCl).
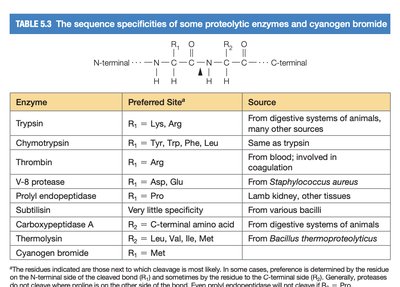
Site-Specific Proteolytic Enzymes
Proteolytic enzymes (proteases) cleave peptide bonds at specific sites, depending on the amino acid sequence. This specificity is crucial for protein processing and degradation.
Examples include trypsin, chymotrypsin, thrombin, and others.
Each enzyme recognizes particular amino acid residues at the cleavage site.
Enzyme | Preferred Site | Source |
|---|---|---|
Trypsin | Lys, Arg | Digestive systems of animals |
Chymotrypsin | Tyr, Trp, Phe, Leu | Digestive systems of animals |
Thrombin | Arg | Blood |
V-8 protease | Asp, Glu | Staphylococcus aureus |
Prolyl endopeptidase | Pro | Lamb kidney, other tissues |
Subtilisin | Very little specificity | Bacilli |
Carboxypeptidase A | C-terminal amino acid | Digestive systems of animals |
Thermolysin | Leu, Val, Ile, Met | Bacillus thermoproteolyticus |
Cyanogen bromide | Met | Chemical reagent |
Peptides and Polypeptides
Peptides are short chains of amino acids, while polypeptides with more than 50 residues are generally referred to as proteins. Each peptide has an unreacted amino group (N-terminus) and an unreacted carboxylic acid group (C-terminus).
The sequence of a polypeptide is written from the N-terminal to the C-terminal end.
Sequences can be represented using three-letter or one-letter abbreviations for amino acids.
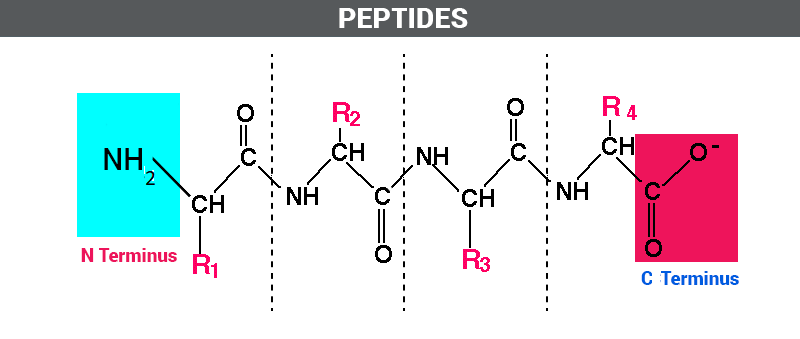
Writing and Interpreting Polypeptide Sequences
Polypeptide sequences are written with the N-terminal residue on the left and the C-terminal residue on the right. Abbreviations help simplify sequence notation.
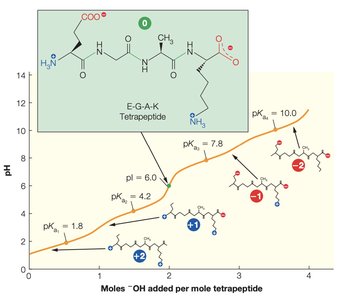
Polyampholytes and Ionization of Side Chains
Proteins are polyampholytes, meaning their amino acid side chains can display a range of pKa values due to differences in the local electrostatic environment. Changes in pH significantly affect the stability and functional properties of peptides.
Side chains may be ionized or neutral depending on the pH.
Ionization affects protein folding, activity, and interactions.
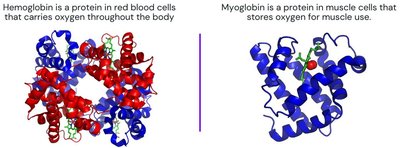
Polypeptide Examples: Hemoglobin and Myoglobin
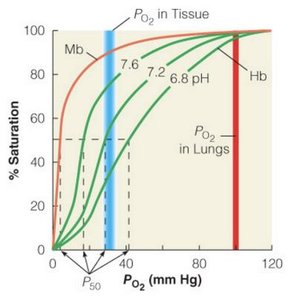
Hemoglobin and myoglobin are classic examples of protein function. Hemoglobin carries oxygen in red blood cells, while myoglobin stores oxygen in muscle cells. Their structures and oxygen-binding properties illustrate the diversity and specificity of protein function.
The Bohr Effect
The Bohr effect describes how changes in pH and CO2 concentration affect hemoglobin's affinity for oxygen. Lower pH (higher acidity) decreases affinity, facilitating oxygen release in tissues.

Primary Structure of Proteins
The primary structure of a protein is defined by the specific number and order of amino acid residues in its polypeptide chain(s). This sequence is unique to each protein and species, and determines higher-order structure and function.
Proteins may consist of one or more polypeptide chains.
Interactions between chains can be noncovalent or covalent.
The sequence is encoded by the organism's genetic material.
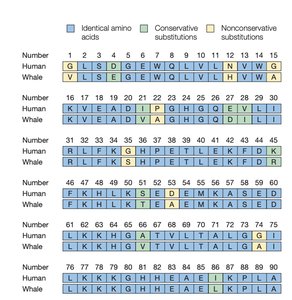
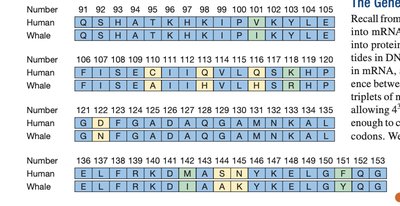
Sequence Comparison: Human vs. Whale
Comparing protein sequences across species reveals evolutionary relationships and functional conservation. Identical, conservative, and nonconservative substitutions can be identified in sequence alignments.
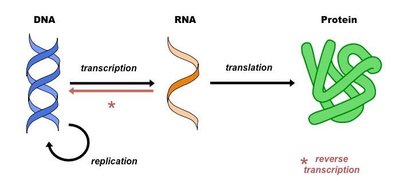
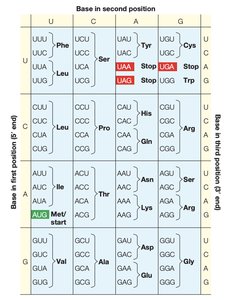
Gene to Protein: The Genetic Code
Proteins are synthesized according to the genetic code, which consists of codons—triplets of nucleotides in DNA or RNA that specify each amino acid. The process involves transcription (DNA to RNA) and translation (RNA to protein).
Each codon corresponds to a specific amino acid or a stop signal.
The genetic code is nearly universal among organisms.


Additional info: The notes above expand on the original content by providing definitions, context, and examples for each topic, ensuring completeness and academic quality for biochemistry students.
