 Back
BackIntroduction to Proteins: The Primary Level of Protein Structure
Study Guide - Smart Notes

Introduction to Proteins: The Primary Level of Protein Structure
Molecules with Multiple Ionizing Groups
Many biologically relevant molecules, such as amino acids, contain both acidic and basic groups, making them ampholytes. These molecules exhibit complex behavior during titration due to the presence of multiple ionizing groups.
Ampholyte: A molecule with both acidic and basic pKa values.
Example: Glycine (H2N-CH2-COOH) is an ampholyte.
Titration Steps: Glycine titration occurs in two steps: first, the more acidic α-carboxylate group is deprotonated, followed by the less acidic α-amine group.
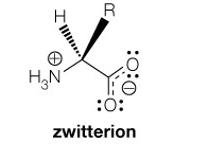
Net Charge at pH 6: At pH 6, glycine exists as a zwitterion, with a net charge of zero.
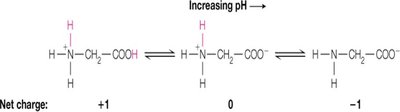
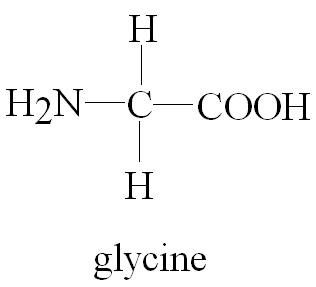
Additional info: Zwitterions are molecules or ions having separate positively and negatively charged groups.
Isoelectric Point (pI)
The isoelectric point is the pH at which the average charge on the molecule is zero. For simple molecules, it is calculated as the average of the two pKa values surrounding the isoelectric species.
Formula:
Example (Glycine):
Example (Aspartic Acid): Aspartic acid has three ionizable groups; pI is calculated by averaging the two pKa values describing the ionization of the isoelectric species:
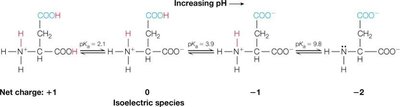
Macroions and Electrostatic Interactions
Macroions are large molecules such as polyelectrolytes (nucleic acids) and polyampholytes (proteins) that carry substantial net charge in solution. Their behavior is influenced by electrostatic forces.
Electrostatic Repulsion: Macroions of like charges repel each other, increasing solubility (e.g., DNA).
Electrostatic Attraction: Oppositely charged macroions associate, as seen in protein-DNA interactions.
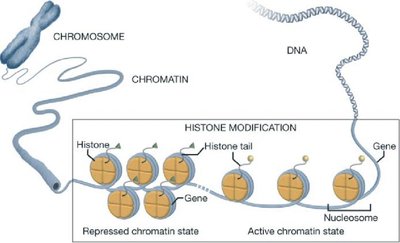
Example: Histones (positively charged proteins) strongly associate with negatively charged DNA to form chromatin.
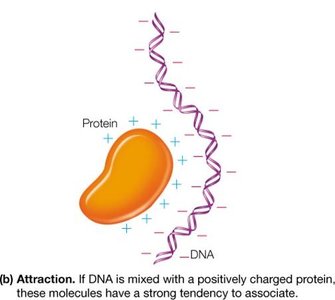
5.1 Amino Acids
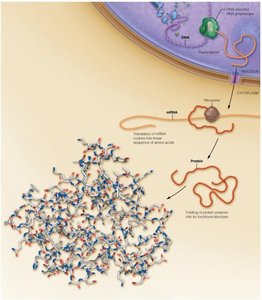
Overview of Protein Synthesis in Eukaryotic Cells
Proteins are synthesized through a process involving transcription and translation. DNA is transcribed to mRNA in the nucleus, which is then exported to the cytoplasm and translated into a linear sequence of amino acids that folds into a three-dimensional structure.
Function of Proteins: Transport, structural framework, muscle contraction, immune responses, blood clotting, and catalysis (enzymes).
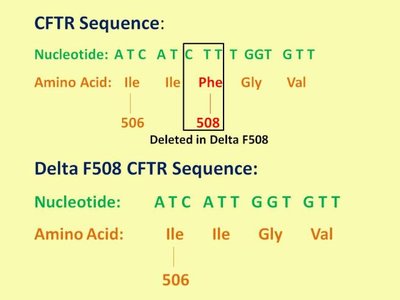
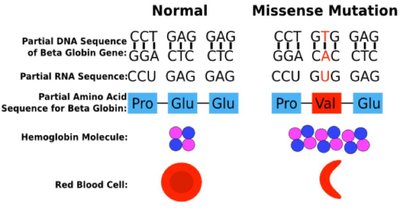
Structure-Function Relationship: The structure of a protein is directly related to its function. Even modest changes in protein structure can lead to instability or loss of function, causing diseases such as cystic fibrosis and sickle-cell anemia.


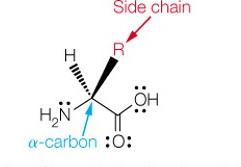
The Structure of α-Amino Acids
Each α-amino acid consists of a central α-carbon attached to an amino group, a carboxylic acid group, a hydrogen atom, and a unique side chain (R group). The properties of amino acids are determined by their side chains.
At physiological pH (~7): The carboxylic acid group is deprotonated (COO-), and the amino group is protonated (NH3+), forming a zwitterion.
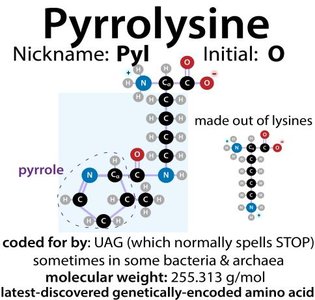
There are 20 common amino acids incorporated into proteins during translation, plus two rare genetically encoded amino acids: selenocysteine and pyrrolysine.

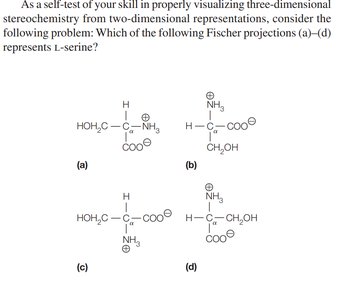
Stereochemistry of the α-Amino Acids
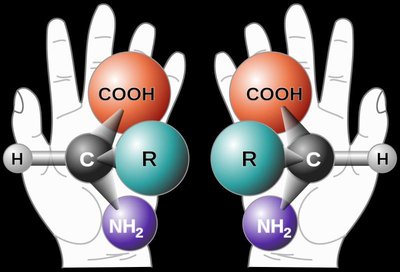
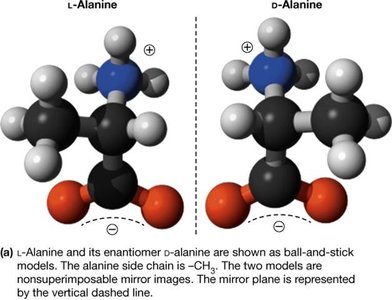
Most amino acids are chiral, meaning their α-carbon is attached to four different groups, resulting in two enantiomers (L and D forms). Proteins are synthesized almost exclusively from L-amino acids.
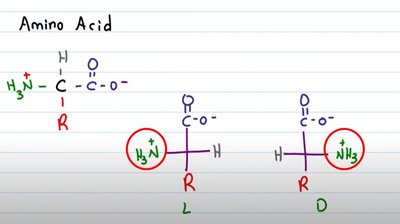
Chirality: Non-superimposable mirror images (enantiomers).
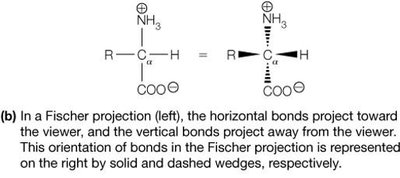
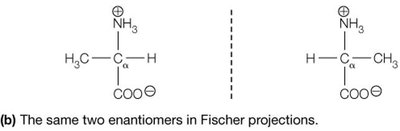
Fischer Projection: A 2D representation of stereochemistry; horizontal bonds project forward, vertical bonds project backward.
Biological Preference: Nearly all naturally occurring proteins use L-amino acids, which is essential for proper protein function and molecular recognition.
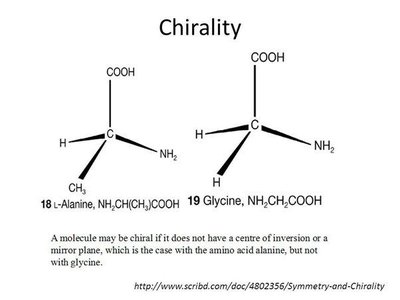
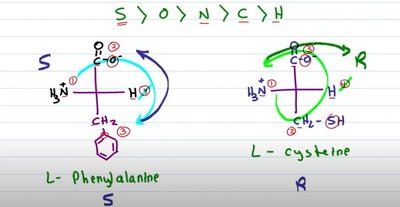
Exceptions: Glycine is not chiral; cysteine is the only R amino acid in biological systems.
Classification of Naturally Occurring Amino Acids
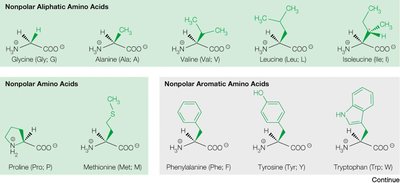
The 20 common amino acids are classified based on the chemical properties of their side chains:
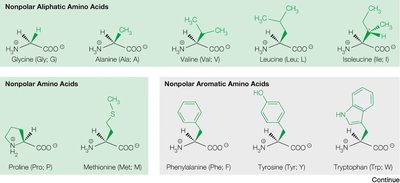
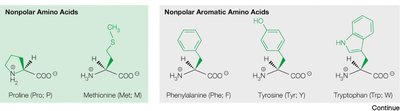
Non-polar aliphatic (e.g., glycine, alanine, valine, leucine, isoleucine)
Non-polar aromatic (e.g., phenylalanine, tyrosine, tryptophan)
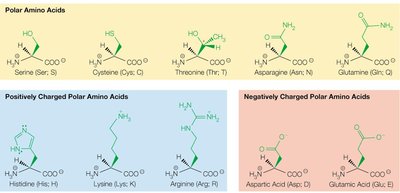
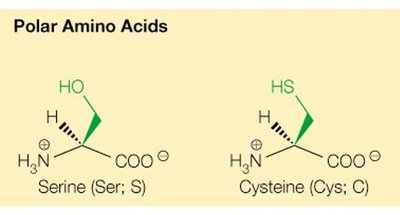
Polar uncharged (e.g., serine, threonine, cysteine, asparagine, glutamine)
Positively charged (basic) (e.g., lysine, arginine, histidine)
Negatively charged (acidic) (e.g., aspartic acid, glutamic acid)
General Properties of Amino Acids
Amino acids exhibit a wide variety of chemical groups in their side chains, allowing proteins to have diverse structures and properties. The side chains determine the hydrophobicity, charge, and reactivity of each amino acid.
Hydrophobic amino acids: Usually found in the core of folded proteins.
Hydrophilic amino acids: Often found on the surface of proteins, interacting with the aqueous environment.
Special roles: Cysteine forms disulfide bonds; serine and cysteine are good nucleophiles and play key roles in enzyme activity.
5.2 Peptides and the Peptide Bond
Peptide Bond Formation
Amino acids are covalently linked by peptide bonds (amide bonds) formed between the α-carboxyl group of one amino acid and the α-amino group of another. Peptide bond formation is a dehydration synthesis reaction, requiring ATP input.
Peptide bond: Amide bond linking amino acids.
Dipeptide: Two amino acids joined by a peptide bond.
Energetics: Peptide formation is thermodynamically unfavorable but kinetically stable due to high activation energy.
Structure of the Peptide Bond
The peptide bond has unique structural properties:
Amide carbonyl (C=O) and amide N-H bonds are nearly parallel.
Six atoms (CCONHC) are coplanar.
Limited rotation due to partial double-bond character.
Peptide bond exists as a resonance hybrid.
Trans configuration is favored, except for proline, which can adopt cis configuration.
Peptide Bond Cleavage
Peptide bonds can be cleaved by hydrolysis, which is thermodynamically favored but kinetically slow without catalysis. Proteolytic enzymes (proteases) and chemical agents (e.g., cyanogen bromide) can cleave specific peptide bonds.
Proteases: Enzymes that cleave specific peptide bonds.
Cyanogen bromide: Cleaves amide bond on C-terminal side of methionine residue.
Oligopeptides and Polypeptides
Oligopeptides are short peptide chains (3–15 residues), polypeptides contain more than 15 residues, and proteins contain more than 50 residues. The sequence of amino acids determines the structure and function of the protein.
Peptides and Proteins as Polyampholytes
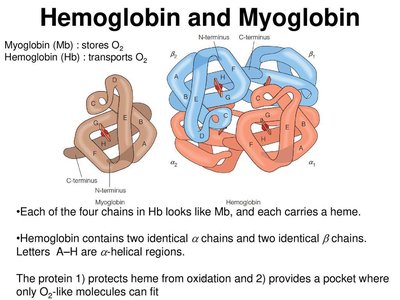
Proteins can act as polyampholytes, carrying both positive and negative charges. This property is important in physiological processes such as the Bohr effect in hemoglobin, which facilitates oxygen release in tissues with high CO2 and low pH.
5.4 From Gene to Protein
The Genetic Code
DNA sequences are transcribed into mRNA, which is translated into proteins. The genetic code uses triplets of nucleotides (codons) to specify amino acids. Most amino acids have multiple codons, and three codons serve as stop signals.
Start codon: AUG (methionine)
Stop codons: UAA, UAG, UGA
Rare amino acids: UGA codes for selenocysteine, UAG for pyrrolysine in special cases.
Post-Translational Processing of Proteins
After translation, polypeptides undergo folding and post-translational modifications, such as disulfide bond formation and proteolytic cleavage. These modifications are essential for protein function and regulation.
Example: Insulin is synthesized as preproinsulin, processed to proinsulin, and then to active insulin by proteolytic cleavage and disulfide bond formation.
Physiological advantage: Inactive precursors can be stored and rapidly activated when needed.
5.5 From Gene Sequence to Protein Function
Protein Sequence Homology and Evolution
Proteins have defined sequences of amino acids. Sequence similarity between proteins from different species reflects evolutionary relationships. Conservative mutations preserve chemical properties, while non-conservative mutations can have significant effects on protein function.
Sequence identity: Exact match of amino acids.
Sequence similarity: Based on chemical properties of side chains.
Structural context: The effect of amino acid substitution depends on whether the site is solvent-exposed or buried in the protein core.
Alignments of Proteins—Globins
Sequence alignments are used to compare protein sequences and infer evolutionary relationships. Identity refers to exact matches, while similarity considers chemical properties.
