 Back
BackPrimary Protein Structure and Sequencing: Study Notes for Biochemistry
Study Guide - Smart Notes

Primary Protein Structure
Introduction to Protein Structure
The structure and function of proteins are dictated by their sequence of amino acids, known as the primary structure. Understanding this sequence is fundamental for biochemists, as it underpins protein folding, function, and interactions. Misfolding or mutations in the primary structure can lead to diseases and altered biological activity.
Primary structure: Linear sequence of amino acids from N-terminus to C-terminus.
Secondary structure: Local spatial arrangement (e.g., α-helix, β-sheet).
Tertiary structure: Three-dimensional folding of a single polypeptide.
Quaternary structure: Association of multiple polypeptide chains.
Function is dictated by protein structure!
Examples of Protein Structure Levels
Each level of protein structure contributes to the overall function and stability of the protein. For example, insulin consists of two chains (A and B) connected by disulfide bonds, illustrating both primary and quaternary structure.

Protein Sequencing
Importance of Protein Sequencing
Protein sequencing reveals the exact order of amino acids, enabling the study of gene-to-protein relationships, drug design, enzyme mechanisms, protein engineering, evolutionary biology, and analytical techniques. Sequencing is foundational for understanding protein function and dysfunction.
Gene-to-protein relationship: Links genomics to proteomics.
Drug design: Targets specific residues or motifs.
Enzyme mechanisms: Identifies catalytic residues.
Protein engineering: Enables rational design of new proteins.
Evolutionary insight: Sequence comparison reveals homology.
Steps in Protein Sequencing
Sequencing a protein involves several key steps:
Purification: Isolate the protein.
Determine number of polypeptides: Analyze N- and C-termini.
Cleavage of disulfide bonds: Separate chains and prevent refolding.
Amino acid composition: Hydrolyze and analyze liberated amino acids.
Fragmentation: Use endopeptidases or chemical reagents to break protein into smaller peptides.
Sequencing fragments: Sequence each fragment and align overlapping regions.
End Group Analysis
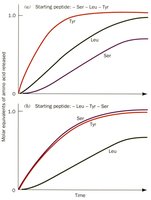
Determining the number of peptides in a protein is achieved by analyzing the N- and C-termini. Dansyl chloride reacts with amines at the N-terminus and lysine side chains, allowing identification of terminal residues.
Amino Acid Composition Analysis
The amino acid composition is determined by complete hydrolysis of the protein, followed by quantitative analysis. Acid hydrolysis destroys some amino acids, while base hydrolysis affects others. An amino acid analyzer is used to quantitate residues after derivatization.
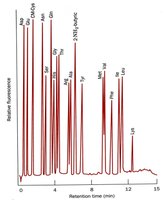
Common amino acids: Leu, Ala, Gly, Ser, Val, Glu, Ile.
Rare amino acids: His, Met, Cys, Trp.
Structural proteins: Often have repeating sequences (e.g., collagen).
Fragmentation of Proteins
Proteins are fragmented using specific endopeptidases (e.g., trypsin, chymotrypsin) or chemical reagents (e.g., cyanogen bromide). Trypsin cleaves after positively charged residues (R, K), while chymotrypsin targets bulky hydrophobic residues (F, W, Y). Cyanogen bromide cleaves after methionine (M).
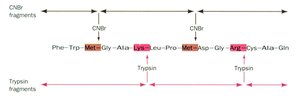
Assembly of Protein Sequence
To assemble the complete sequence, fragments generated by different cleavage methods are aligned based on overlapping regions. Logical deduction and double-checking are essential to ensure accuracy.
Protein Mass Spectrometry
Mass Spectrometry Techniques
Mass spectrometry (MS) is a powerful tool for sequencing proteins. Proteins are digested into fragments (often by trypsin), which are then analyzed by electrospray ionization or MALDI-TOF. The mass/charge (m/z) ratio of fragments is measured, and tandem MS can further fragment ions for sequence determination.
Electrospray: Produces multiply charged ions for MS analysis.
MALDI-TOF: Uses laser ionization and time-of-flight measurement.
Tandem MS: Provides sequence tags for database identification.
Peptide Sequence Tag Notation
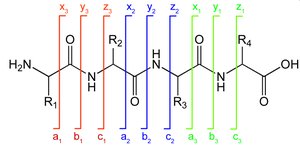
Peptide fragments are labeled based on where the charge is retained: a, b, c (N-terminus) and x, y, z (C-terminus). Subscripts indicate the number of residues, and superscripts denote neutral losses (e.g., * for ammonia, ° for water).
Protein Databases
Protein Database Utility
Protein sequences are stored in databases (e.g., UniProt) for comparison, identification, and study. Each protein entry includes an accession number, organism, known mutations, and modified amino acids. Database searching is essential for modern proteomics and bioinformatics.
Accession number: Unique identifier for each protein.
Sequence comparison: Reveals homology and functional domains.
Mutation tracking: Links sequence changes to phenotypic effects.
