 Back
BackProtein Families: Structure, Evolution, and Functional Implications
Study Guide - Smart Notes

Protein Families
Definition and Importance
Protein families are groups of evolutionarily related proteins that share similar sequences, structures, and often functions. Understanding protein families is fundamental in biochemistry for predicting protein function, studying evolutionary relationships, and informing drug design and disease research.
Definition: A protein family consists of proteins with significant sequence and/or structural similarity, often arising from a common ancestral gene.
Functional Prediction: Membership in a protein family allows inference of function for newly discovered proteins based on known family characteristics.
Evolutionary Insight: Protein families arise through gene duplication and divergence, providing a record of evolutionary history.
Medical Relevance: Many diseases are linked to mutations in specific protein families, making them targets for drug design.
Structural Biology: Conserved 3D folds within families aid in modeling unknown structures and predicting binding sites.
Bioinformatics: Tools like BLAST and Pfam use family information for genome annotation and functional prediction.
Protein Mutation: Sickle Cell Anemia
Example of Protein Mutation
Sickle cell anemia is a classic example of how a single amino acid substitution in a protein can lead to significant physiological consequences. The disease is caused by a mutation in the β-globin chain of hemoglobin, resulting in the replacement of glutamic acid with valine at position 6.
Peptide Mapping: Proteins are digested and separated using two-dimensional chromatography to identify sequence differences.
Sequence Comparison: Normal hemoglobin (HbA): V-H-L-T-P-E-E-K; Sickle cell hemoglobin (HbS): V-H-L-T-P-V-E-K.
Functional Impact: The mutation causes hemoglobin to aggregate under low oxygen, deforming red blood cells into a sickle shape.
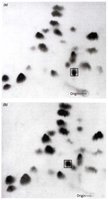
Clinical Manifestation: Sickle-shaped cells are less flexible, leading to blockages in blood vessels and various complications.
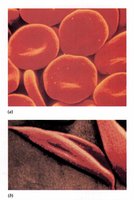
Genetic and Evolutionary Context: The sickle cell trait confers resistance to malaria in heterozygotes, explaining its persistence in certain populations.
Example: Individuals with one sickle cell allele (heterozygotes) are resistant to malaria, while homozygotes for the sickle allele develop sickle cell disease.
Protein Family Example: Cytochrome C
Structure and Function
Cytochrome c is a small hemeprotein involved in the electron transport chain and apoptosis. It is highly conserved across species, making it a model for studying protein evolution.
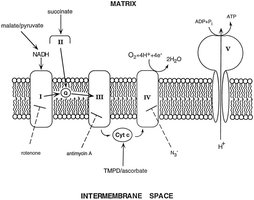
Location: Found in the inner mitochondrial membrane.
Function: Transfers electrons between Complex III and Complex IV in the electron transport chain; also involved in programmed cell death (apoptosis).
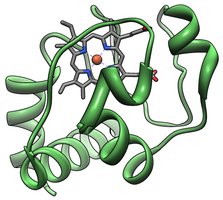
Conservation: The primary structure of cytochrome c is highly conserved, with invariant residues reflecting essential chemical functions.
Types of Residue Changes:
Conserved residues: Invariant, essential for function.
Conserved substitutions: Chemically similar replacements (e.g., Asp for Glu).
Variable regions: Less critical for function, more tolerant to change.
Evolutionary Analysis
Comparing cytochrome c sequences across species reveals evolutionary relationships. The number of amino acid differences can be used to construct phylogenetic trees and estimate evolutionary distances.
Difference Matrix: Quantifies amino acid differences between species.
Phylogenetic Tree: Branch points represent common ancestors; distances reflect sequence divergence.
PAM Units: Percentage of Accepted Mutations, a measure of evolutionary distance.
Example: The difference matrix for cytochrome c among 26 species shows that closely related species (e.g., humans and chimpanzees) have few differences, while distant species (e.g., humans and yeast) have many.
Mutation Rate: Although DNA mutates at a relatively constant rate, some proteins are less tolerant to mutations due to functional constraints.
Time Dependence: Mutation rates appear to be more dependent on time than on the number of generations.
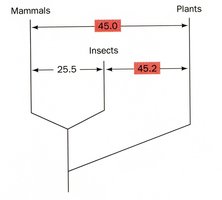
Gene Duplication and Protein Families
Mechanism and Evolutionary Significance
Gene duplication is a major evolutionary mechanism generating new genetic material. Duplicated genes can diverge, leading to the formation of protein families with related but distinct functions.
Definition: Gene duplication is the process by which a region of DNA containing a gene is duplicated, resulting in two or more copies.
Mechanisms: Errors in DNA replication, repair, or chromosomal rearrangements.
Evolutionary Outcome: Duplicated genes can accumulate mutations, leading to new functions or properties.
Example: The globin family (hemoglobin and myoglobin) arose through gene duplication and subsequent divergence.
The Globin Family: Hemoglobin and Myoglobin
The globin family illustrates how gene duplication and divergence can produce proteins with related but specialized functions.
Hemoglobin: Oxygen transport protein in red blood cells; must bind and release oxygen as needed.
Myoglobin: Oxygen storage protein in muscle; binds oxygen tightly and releases it when concentrations are very low.
Evolutionary History:
Primordial globin gene functioned as an oxygen-storage protein.
Duplication and divergence led to specialized forms (e.g., tetrameric hemoglobin, monomeric myoglobin).
Further duplications produced fetal and embryonic hemoglobins with distinct properties.
Protein Evolution vs. Organismal Evolution
Comparative Genomics
Protein sequence similarity does not always correlate directly with organismal differences. For example, humans and chimpanzees share about 99% of their protein sequences, yet exhibit significant phenotypic differences, often due to changes in gene regulation rather than protein sequence.
Key Point: Rapid divergence in phenotype can occur with few changes in protein sequence, often through altered gene expression patterns.
Summary Table: Types of Amino Acid Changes in Protein Families
Type of Change | Description | Example |
|---|---|---|
Invariant (Conserved) Residue | Essential for function; unchanged across species | Active site histidine in cytochrome c |
Conserved Substitution | Replacement with similar chemical property | Asp for Glu |
Variable Region | Not essential for function; tolerant to change | Surface loops |
