 Back
BackProtein Function and Ligand Binding: Oxygen Transport, Regulation, and Immune Response
Study Guide - Smart Notes

Protein Function
Principles of Protein-Ligand Interaction
Proteins perform their functions by interacting dynamically with other molecules, often through reversible binding. These interactions are fundamental to biological processes, including catalysis, signaling, and transport.
Ligand: A molecule reversibly bound by a protein. Ligands can be small molecules, ions, or other proteins.
Binding Site: The region of the protein complementary to the ligand in size, shape, charge, and hydrophobic/hydrophilic character.
Specificity: Proteins selectively bind specific ligands, maintaining cellular order.
Flexibility: Proteins can undergo subtle or dramatic conformational changes, essential for function.
Induced Fit: Ligand binding often induces a conformational change, increasing binding affinity.
Multisubunit Effects: In multisubunit proteins, conformational changes in one subunit can affect others.
Regulation: Protein-ligand interactions may be regulated by allosteric mechanisms.

Oxygen-Binding Proteins
Heme Prosthetic Group and Oxygen Binding
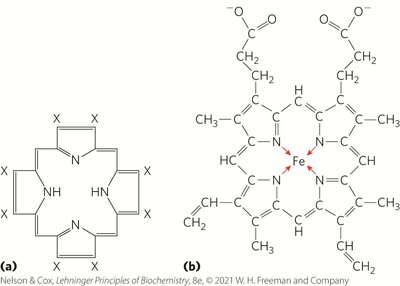
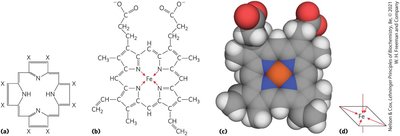
Oxygen transport and storage in vertebrates rely on proteins containing a heme prosthetic group, which binds oxygen reversibly. Heme consists of a protoporphyrin ring with a central Fe2+ ion.
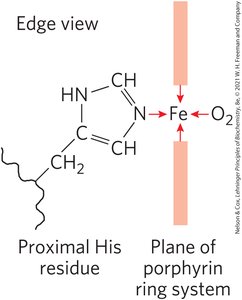
Heme Structure: Four nitrogen atoms in the porphyrin ring coordinate Fe2+, with two additional perpendicular coordination sites.
Oxygen Binding: One perpendicular site binds a histidine residue (proximal His), the other binds O2. Fe2+ binds O2 reversibly; Fe3+ does not.
Globin Family: Myoglobin and Hemoglobin
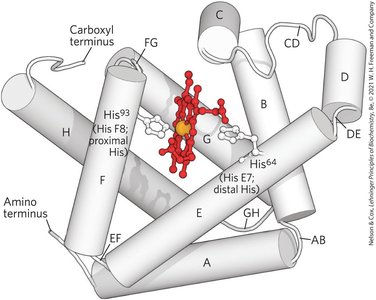
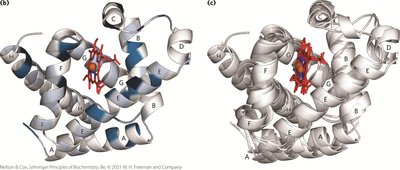
Globins are a widespread protein family with a highly conserved tertiary structure. Myoglobin and hemoglobin are the primary oxygen-binding globins in mammals.
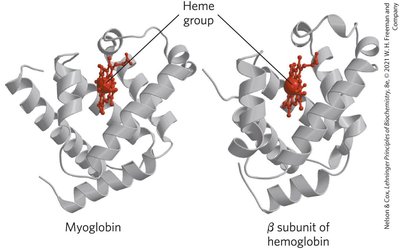
Myoglobin: Monomeric, facilitates O2 diffusion in muscle tissue, contains a single heme group.
Hemoglobin: Tetrameric, transports O2 in blood, consists of two α and two β chains, each with a heme group.
Quantitative Description of Protein-Ligand Binding
The reversible binding of a protein (P) to a ligand (L) is described by equilibrium expressions and constants.
Association Constant (Ka): Measures affinity of ligand for protein. Higher Ka = higher affinity.
Dissociation Constant (Kd): Reciprocal of Ka. Lower Kd = higher affinity.
Fractional Occupancy (Y): Fraction of binding sites occupied by ligand.
Key Equations:
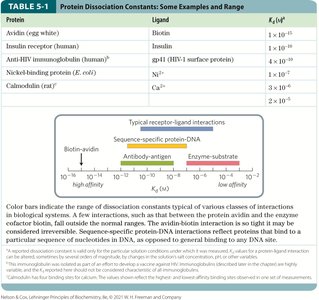
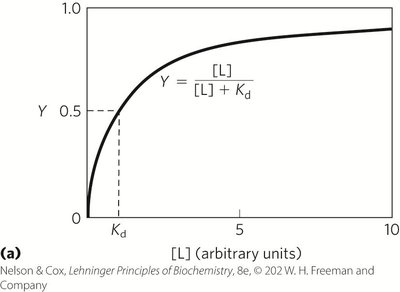

Protein | Ligand | Kd (M) |
|---|---|---|
Avidin (egg white) | Biotin | 1 × 10-15 |
Insulin receptor (human) | Insulin | 1 × 10-9 |
Anti-HIV immunoglobulin (human) | gp41 (HIV-1 surface protein) | 1 × 10-10 |
Nickel-binding protein (E. coli) | Ni2+ | 3 × 10-7 |
Calmodulin (rat) | Ca2+ | 1 × 10-6 |
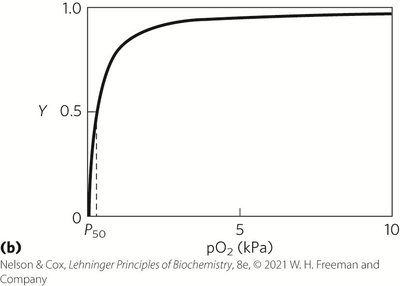
Partial Pressure and Oxygen Binding
Oxygen binding is often described in terms of partial pressure (pO2), with P50 representing the pO2 at which half the binding sites are occupied.
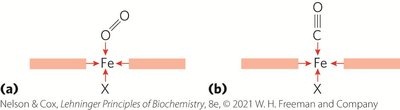
Protein Structure and Ligand Binding
Protein conformation affects ligand binding. For example, carbon monoxide (CO) binds free heme much more strongly than O2 due to differences in orbital structure and binding geometry.
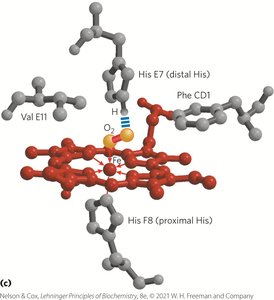

Hemoglobin: Structure and Cooperative Binding
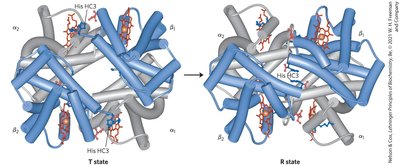
Hemoglobin exhibits cooperative binding, where the binding of O2 to one subunit increases the affinity of other subunits. This is reflected in its sigmoidal binding curve.
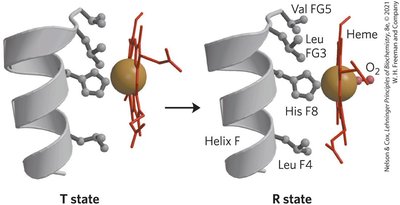
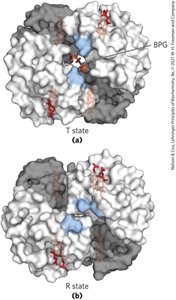
T state: Low affinity for O2, stabilized by ion pairs.
R state: High affinity for O2, favored upon O2 binding.
Cooperativity: Conformational changes in one subunit affect others.
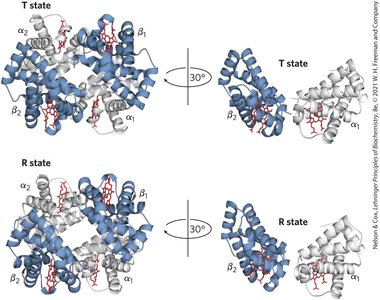
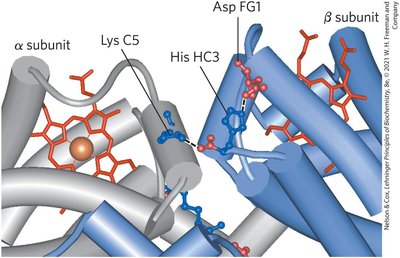

Quantitative Description of Cooperative Binding
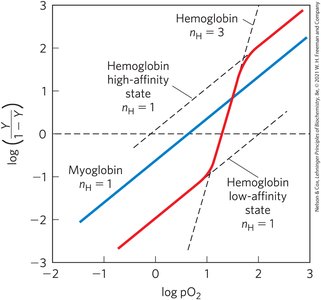
Cooperative binding is described by the Hill equation, which relates ligand concentration to fractional occupancy.
Hill coefficient (nH): Indicates degree of cooperativity. nH = 1 (non-cooperative), nH > 1 (positive cooperativity), nH < 1 (negative cooperativity).
Models of Cooperative Binding
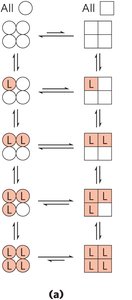
Two models explain cooperative binding in hemoglobin:
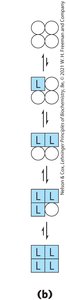
MWC (Concerted) Model: All subunits are in the same conformation; ligand binding shifts equilibrium toward R state.
Sequential Model: Each subunit can change conformation independently; ligand binding progressively favors R state.
Regulation of Oxygen Binding: Bohr Effect and BPG
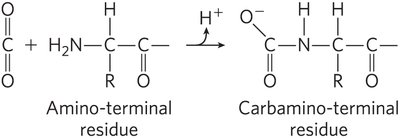
Hemoglobin's affinity for O2 is regulated by pH, CO2, and 2,3-bisphosphoglycerate (BPG).
Bohr Effect: Lower pH and higher CO2 favor the T state, promoting O2 release.
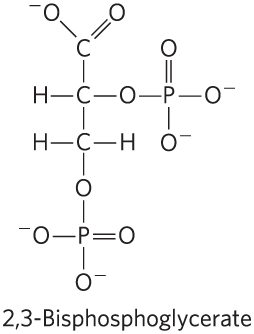
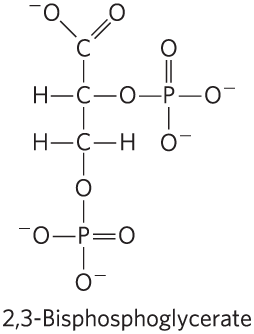
BPG: Binds to hemoglobin, stabilizes T state, reduces O2 affinity, and increases at high altitudes.

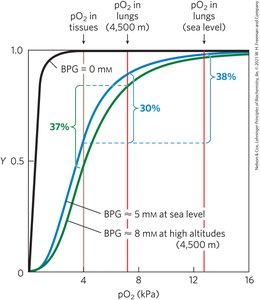
Hemoglobin Variants and Disease
Fetal hemoglobin (α2γ2) has higher O2 affinity due to lower BPG binding. Sickle cell anemia results from a single amino acid substitution, causing hemoglobin polymerization and cell deformation.
Immune System and Immunoglobulins
Immune Response and Antibody Structure
The immune system distinguishes self from nonself and eliminates pathogens. Antibodies (immunoglobulins) are proteins that bind antigens with high specificity and affinity.
Antigen: Molecule capable of eliciting an immune response.
Antibody: Protein produced by B cells, binds antigens at specific sites (epitopes).
Immunoglobulin G (IgG): Major class, consists of two heavy and two light chains, each with variable and constant domains.
Induced Fit: Antibody-antigen binding involves conformational changes for tight interaction.
Antibody-Antigen Interaction and Analytical Applications
Antibodies bind antigens tightly and specifically, forming the basis for analytical techniques such as Western blotting.
Polyclonal Antibodies: Mixture recognizing different parts of a protein.
Monoclonal Antibodies: Homogeneous, recognize a single epitope.
Muscle Contraction: Actin, Myosin, and Molecular Motors
Major Proteins of Muscle
Muscle contraction is driven by transient interactions between actin and myosin filaments, powered by ATP hydrolysis.
Myosin: Forms thick filaments, consists of two heavy and four light chains, with a coiled-coil tail and globular head.
Actin: Monomeric G-actin forms F-actin (thin filaments), each monomer binds and hydrolyzes ATP.
Thick Filaments: Aggregated myosin.
Thin Filaments: F-actin, troponin, and tropomyosin.
Muscle Fiber Organization and Sarcomere Structure
Muscle fibers contain myofibrils organized into sarcomeres, the fundamental contractile units.
Sarcomere: Contains thick and thin filaments, defined by A band, I band, Z disk, and M line.
Regulation of Muscle Contraction
Muscle contraction is regulated by tropomyosin and troponin, which control access to myosin-binding sites on actin.
Tropomyosin: Blocks myosin-binding sites on actin.
Troponin: Binds Ca2+, causes conformational change, exposes myosin-binding sites.
Summary: Protein function is governed by dynamic, reversible interactions with ligands, regulated by structural changes, allosteric effects, and environmental factors. These principles underpin oxygen transport, immune response, and muscle contraction.
