 Back
BackProtein Secondary Structure: Foundations, Helices, Sheets, and Loops
Study Guide - Smart Notes

Protein Secondary Structure
Introduction to Protein Secondary Structure
The secondary structure of proteins refers to the local spatial arrangement of the polypeptide backbone, stabilized primarily by hydrogen bonding. Understanding secondary structure is essential for biochemistry students, as it forms the basis for protein folding, function, and interactions in biological systems. The main types of secondary structure are alpha helices, beta sheets, and loops/turns.
Structure Dictates Function: The 3D shape of proteins, determined by secondary structure, is crucial for their biological activity.
Protein Folding and Stability: Secondary structure elements stabilize proteins via hydrogen bonds and other weak interactions.
Basis for Advanced Topics: Knowledge of secondary structure is foundational for understanding tertiary/quaternary structure, enzyme mechanics, and protein engineering.
Health and Disease: Misfolding of secondary structures is linked to diseases such as Alzheimer's and prion diseases.
Experimental Relevance: Techniques like X-ray crystallography and circular dichroism analyze secondary structure.
Bioinformatics: Secondary structure prediction aids in modeling unknown proteins.
Three-Dimensional Protein Structures
Proteins adopt specific three-dimensional conformations, known as their native state, stabilized by weak interactions:
Hydrophobic interactions
Hydrogen bonds
Ionic bonds
Van der Waals attractions
A denatured protein loses its native conformation, often resulting in loss of function and precipitation.
The Amide (Peptide) Bond
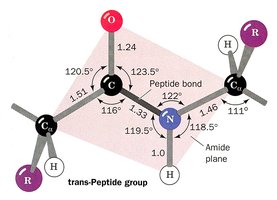
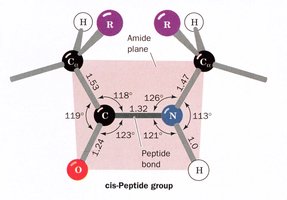
The peptide bond, formed between the carboxyl group of one amino acid and the amino group of another, is planar due to partial double-bond character from resonance. Most peptide bonds are in the trans configuration, but those preceding proline may be cis.
Bond Lengths and Angles: The C-N peptide bond is shorter than a typical C-N bond due to resonance.
Planarity: Peptide bonds are planar, maximizing resonance stabilization.
Cis vs. Trans: Trans is favored except for proline, where cis is more common.
Torsion Angles: Phi (\(\phi\)) and Psi (\(\psi\))
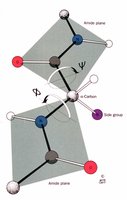
The backbone conformation of proteins is described by two key dihedral angles:
Phi (\(\phi\)): Rotation around the N-C\(\alpha\) bond
Psi (\(\psi\)): Rotation around the C\(\alpha\)-C bond
When the peptide chain is fully extended, both angles are ±180°. These angles determine the overall backbone structure.
Ramachandran Plot
The Ramachandran plot is a graphical representation of the allowed regions for phi and psi angles in a polypeptide. It shows which conformations are sterically possible, based on backbone geometry and side chain interactions.
X-axis: Phi (\(\phi\)) angle
Y-axis: Psi (\(\psi\)) angle
Allowed regions: Correspond to common secondary structures (alpha helix, beta sheet, etc.)
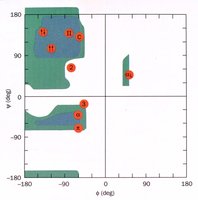
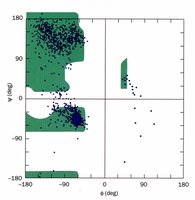
Secondary Structure Types and Their Torsion Angles
Secondary structure elements are defined by characteristic phi and psi angles:
Alpha helix (right-handed): \(\phi = -57^\circ\), \(\psi = -47^\circ\)
Beta sheet: \(\phi = -119^\circ\), \(\psi = 113^\circ\)
310 helix: \(\phi = -49^\circ\), \(\psi = -26^\circ\)
Collagen helix: \(\phi = -51^\circ\), \(\psi = 153^\circ\)
Helices
Alpha Helix
The alpha helix is the most common helical structure in proteins, characterized by a right-handed spiral stabilized by hydrogen bonds.
Residues per turn: 3.6
Pitch: 5.4 Å
Hydrogen bonding: C=O of residue n bonds to N-H of residue n+4
Bond length: Hydrogen bond is 2.8 Å, nearly optimal
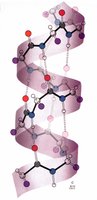
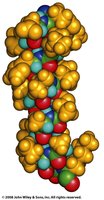
Helix Nomenclature and Variants
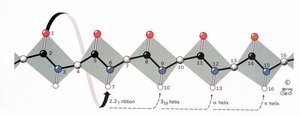
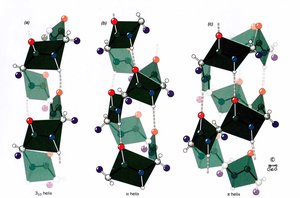
Helices are classified by the number of residues per turn (N) and the number of atoms in the hydrogen-bonded ring (M):
2.27 Ribbon: N = 2.2, hydrogen bond from atom 1 to atom 7
310 Helix: N = 3, hydrogen bond from atom 1 to atom 10
Pi Helix: N = 4.4, rarely observed
Proline Helix and Collagen
The proline helix is a left-handed helix with 3 residues per turn and a pitch of 9.4 Å. It lacks backbone hydrogen bonding but is stabilized by the unique properties of proline and glycine, commonly found in collagen.
Beta Structures
Beta Sheets
Beta sheets are formed by hydrogen bonding between adjacent peptide chains, resulting in a pleated, extended structure. Sheets can be parallel or antiparallel, with antiparallel sheets being more stable due to optimal hydrogen bonding.
Repeat distance: 7.0 Å
R group orientation: Alternates above and below the sheet
Strands per sheet: 2–15, average 6
Width: 25 Å
Twist: Due to chiral amino acids, sheets exhibit a right-handed twist

Loop Structures
Turns, Coils, and Loops
Non-repetitive regions of proteins, such as turns, coils, and loops, connect secondary structure elements and contribute to protein flexibility and function. About 50% of globular protein structure consists of these regions.
Beta bends: Hairpin turns between antiparallel sheets, classified as Type I and Type II
Type I: \(\phi_2 = -60^\circ\), \(\psi_2 = -30^\circ\); \(\phi_3 = -90^\circ\), \(\psi_3 = 0^\circ\)
Type II: \(\phi_2 = -60^\circ\), \(\psi_2 = 120^\circ\); \(\phi_3 = 90^\circ\), \(\psi_3 = 0^\circ\)
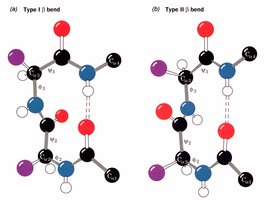
Loops may be disordered and flexible, which is often necessary for protein function. Globular proteins (enzymes) and fibrous proteins (structural) differ in their loop content and function.
Summary Table: Secondary Structure Types
Structure | Phi (\(\phi\)) | Psi (\(\psi\)) | Stabilization | Residues/Turn |
|---|---|---|---|---|
Alpha helix | -57° | -47° | Hydrogen bonds (n to n+4) | 3.6 |
Beta sheet | -119° | 113° | Inter-strand hydrogen bonds | N/A |
310 helix | -49° | -26° | Hydrogen bonds (n to n+3) | 3 |
Collagen helix | -51° | 153° | Proline/Glycine content | 3 |
Conclusion
Understanding protein secondary structure is fundamental for biochemistry students, providing insight into protein folding, stability, function, and disease. Mastery of these concepts is essential for advanced study in structural biology, enzymology, and biotechnology.
