 Back
BackProtein Structure and Sequencing Techniques in Biochemistry
Study Guide - Smart Notes

Protein Structure: Levels and Functions
Overview of Protein Functions
Proteins are essential biomolecules with diverse functions, all of which are closely linked to their structural complexity. Their roles include catalysis (enzymes), transport (hemoglobin, albumin), structural support (spectrin), mechanical work (actin, myosin), regulation (transcription factors), signaling (hormones, receptors), and immunity (immunoglobulins).
Catalysis: Enzymes accelerate biochemical reactions.
Transport: Proteins like hemoglobin carry oxygen; albumin transports various molecules.
Structure: Spectrin maintains cell shape.
Mechanical Work: Actin and myosin enable muscle contraction.
Regulation: Proteins regulate gene expression.
Signaling: Hormones and receptors mediate cellular communication.
Immunity: Immunoglobulins recognize and neutralize pathogens.
Levels of Protein Structure
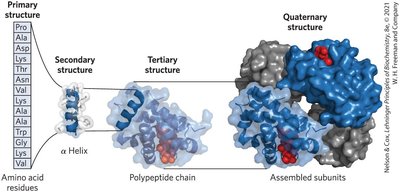
Protein structure is organized into four hierarchical levels, each contributing to the molecule's function and stability.
Primary Structure: The linear sequence of amino acids linked by covalent peptide bonds.
Secondary Structure: Regular structural motifs such as α-helices and β-sheets, stabilized by hydrogen bonds.
Tertiary Structure: The three-dimensional folding of a single polypeptide chain, driven by interactions among side chains.
Quaternary Structure: Assembly of multiple polypeptide subunits into a functional protein complex.
Determination of Primary Structure
Sequencing Methods
Determining the primary structure of proteins is fundamental for understanding their function and for applications in biotechnology and medicine. Two main approaches are used: direct protein sequencing and sequencing of the encoding DNA.
Direct Protein Sequencing: Involves cleavage of proteins into peptides, followed by sequencing using chemical or enzymatic methods.
DNA Sequencing: Sequencing the cDNA encoding the protein provides the amino acid sequence.
Edman Degradation
The Edman degradation method sequentially removes the N-terminal amino acid from a peptide, allowing identification of each residue.
Labeling the N-terminal amino acid with phenylisothiocyanate (PITC).
Selective cleavage to release the labeled amino acid.
Identification of the released amino acid, typically by HPLC.

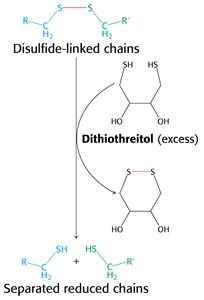
Disulfide Bond Cleavage
Disulfide bonds stabilize protein structure and must be reduced to separate polypeptide chains for sequencing. Dithiothreitol (DTT) is commonly used for this purpose.
Protease Specificity
Proteases are enzymes that cleave peptide bonds at specific amino acid residues, facilitating the generation of peptide fragments for sequencing.
Trypsin: Cleaves after lysine (Lys) or arginine (Arg).
Chymotrypsin: Cleaves after phenylalanine (Phe), tryptophan (Trp), or tyrosine (Tyr).
Other proteases: Have unique specificities, enabling tailored fragmentation.

Peptide Fragment Assembly
After sequencing individual peptide fragments, overlapping sequences are used to reconstruct the full protein sequence.

Mass Spectrometry in Protein Sequencing
Principles of Mass Spectrometry
Mass spectrometry (MS) is a powerful technique for protein identification and sequencing. It measures the mass-to-charge ratio (m/z) of ionized peptides.
Ionization Methods: Electrospray ionization (ESI) and matrix-assisted laser desorption/ionization (MALDI) are commonly used.
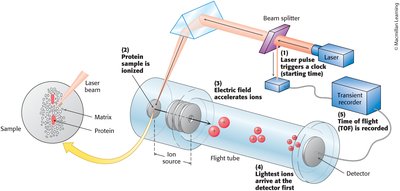
Mass Analyzers: Devices such as Orbitrap and time-of-flight (TOF) separate ions based on m/z.
Detection: The detector records the abundance of ions at each m/z value.

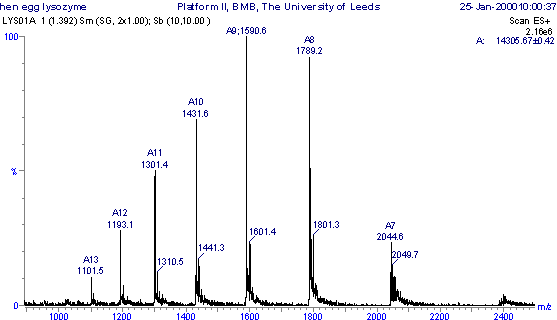
Interpretation of Mass Spectra
Each peak in a mass spectrum corresponds to a peptide ion with a specific m/z value. The charge state and mass of the peptide can be calculated using:
Equation:
n: Number of charges (protons).
H+: Mass of a proton (1.0 Da).

Bottom-Up Proteomics
Proteins are digested into peptides, which are then analyzed by MS to deduce the primary structure and identify the protein.

Tandem Mass Spectrometry (MS/MS)
MS/MS involves fragmentation of peptide ions, allowing determination of the amino acid sequence from the resulting fragment ions.
Isolation: Select a peptide ion.
Fragmentation: Break the peptide into smaller ions.
Analysis: Record the m/z of fragment ions to deduce sequence.

Evolution and Bioinformatics of Proteins
Sequence Variation and Evolution
Protein sequences evolve through mutations, which may be conserved, variable, or confer selective advantages. Sequence comparison reveals evolutionary relationships and functional importance.
Invariant Residues: Essential for function, highly conserved.
Conservative Substitutions: Similar properties, often tolerated.
Hypervariable Positions: Many residues tolerated, less functional constraint.
Sickle Cell Anemia: A Case Study
Sickle cell anemia is caused by a single amino acid substitution (Glu to Val) in the β-chain of hemoglobin, affecting protein function and conferring resistance to malaria in heterozygotes.
Mutation: Glutamate 6 → Valine in β-globin.
Effect: Alters hemoglobin structure, leading to rigid, sickle-shaped red blood cells.
Clinical Implication: Reduced oxygen transport and increased hemolysis.
Protein Homology and Phylogenetics
Homologous proteins share sequence similarity and can be classified as orthologs (between species) or paralogs (within a species). Sequence comparison enables construction of phylogenetic trees and identification of evolutionary relationships.
PAM Units: Percentage of accepted point mutations per 100 residues quantifies evolutionary distance.
Phylogenetic Trees: Visualize relationships and common ancestors.
Gene Duplication and Protein Families
Gene duplication produces paralogous proteins, allowing functional diversification. The globin family exemplifies this process, with hemoglobin and myoglobin evolving distinct roles in oxygen transport and storage.
Hemoglobin: Tetrameric, transports oxygen.
Myoglobin: Monomeric, stores oxygen.
Additional info: These notes expand on the original slides by providing definitions, context, and examples for each major concept, ensuring a comprehensive and self-contained study guide for biochemistry students.
