Back
BackProtein Structure Determination and Folding: Study Notes for Biochemistry
Study Guide - Smart Notes

Protein Structure: Tertiary and Quaternary Levels
Overview of Protein Structure
Proteins possess complex three-dimensional structures that are essential for their biological function. The tertiary structure refers to the overall folding of a single polypeptide chain, while quaternary structure describes the arrangement of multiple polypeptide subunits. Understanding these structures is crucial for interpreting protein function, disease mechanisms, and drug design.
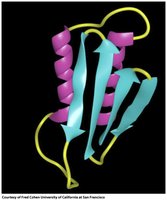
Tertiary Structure: Involves the spatial arrangement of secondary structural elements (α-helices, β-sheets) into motifs and domains.
Quaternary Structure: Describes the assembly of multiple polypeptide chains (subunits) into a functional protein complex.
Conservation: Protein structure is often more conserved than sequence across evolutionary time.
Techniques for Determining Protein Structure
X-ray Crystallography
X-ray crystallography is a powerful method for determining the atomic structure of proteins. It requires the formation of protein crystals, which are then exposed to X-rays to produce diffraction patterns. These patterns are analyzed to generate electron density maps and build atomic models.
Advantages: High resolution (up to 1 Å), can resolve all side chains, suitable for large complexes and membrane proteins.
Disadvantages: Requires crystallization, yields static structures.
Workflow: Purify protein → Crystallize → Collect diffraction data → Solve phase problem → Generate electron density map → Build structure.


Crystallization Methods: Vapor Diffusion
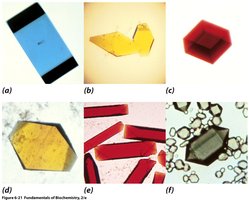
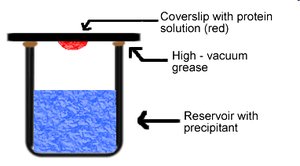
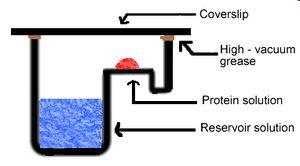
Vapor diffusion is a common technique for protein crystallization. It involves equilibrating a drop of protein solution against a reservoir containing precipitant, allowing gradual concentration changes that promote crystal formation.
Setup: Protein solution is placed on a coverslip above a reservoir; water vapor diffuses, concentrating the protein.
Applications: Used in microtiter plates to screen many conditions.
Bragg's Law and Data Collection
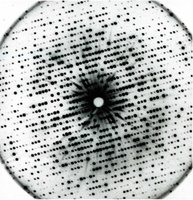
X-ray diffraction produces a pattern of spots, each corresponding to a set of lattice planes in the crystal. The position and intensity of these spots are analyzed using Bragg's Law:
Bragg's Law:
Parameters: = order, = wavelength, = spacing, = angle.
Miller Indices: Spots are indexed by (h, k, l).
Phase Problem in X-ray Crystallography
The phase problem arises because diffraction patterns provide intensity but not phase information, which is essential for constructing electron density maps. Solutions include direct methods, molecular replacement, and isomorphous replacement.

Electron Density Maps and Resolution
Electron density maps are generated from diffraction data and used to build atomic models. Resolution indicates the level of detail; higher resolution allows visualization of individual atoms and even hydrogens.
Cryo-Electron Microscopy (Cryo-EM)
Cryo-EM is a technique that images proteins in a hydrated, glass-like state at cryogenic temperatures. It is especially useful for large complexes and membrane proteins, and does not require crystallization.
Advantages: No phase problem, rapid structure determination, suitable for large molecules.
Disadvantages: Lower resolution for small proteins, slower than crystallography.
Applications: Used to solve structures such as the COVID-19 spike protein.
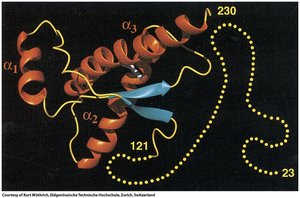
Nuclear Magnetic Resonance (NMR) Spectroscopy
NMR spectroscopy determines protein structures in solution, providing information about dynamic conformations. It relies on isotopic labeling and is best suited for smaller proteins.
Advantages: Proteins are in solution, structures are dynamic.
Disadvantages: Requires specific isotopes (1H, 13C, 15N), limited to smaller proteins (<40 kDa).
Techniques: COSY and NOESY NMR provide information on bond angles and spatial proximity.
Protein Folding Motifs and Families
Folding Motifs
Specific amino acid sequences fold into characteristic motifs, such as β-α-β, β-hairpin, and α-α motifs. These motifs combine to form domains and contribute to the overall tertiary structure.
Examples: β-barrel motifs, Rossmann fold for nucleotide binding.
Protein Families: Proteins with similar folding patterns are grouped into families; structure is often more conserved than sequence.
Quaternary Structure and Symmetry
Subunit Arrangement
Quaternary structure involves the noncovalent association of protein subunits, which may be identical (protomers) or different. Symmetry is a key feature, with cyclic (C2, C3, C6) and dihedral symmetries commonly observed.
Rotational Symmetry: Subunits are arranged around a central axis.
Dihedral Symmetry: Combines rotational and perpendicular two-fold symmetry.
Protein Stability, Folding, and Dynamics
Forces Stabilizing Proteins
Protein stability is governed by electrostatic interactions, hydrogen bonds, van der Waals forces, and hydrophobic effects. Hydrophobic residues tend to be buried in the protein interior, while polar residues are exposed.
Disulfide Bonds: Covalent cross-links between cysteine residues stabilize structure.
Metal Ions: Zinc fingers stabilize DNA binding domains.
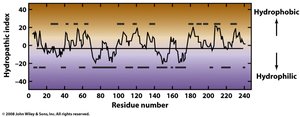
Hydrophobicity Scale
The hydrophobicity scale predicts which amino acid residues are likely to be found in the protein interior or on the surface. Hydropathy plots can be used to analyze protein sequences.
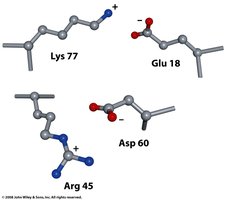
Ion Pairs and Zinc Fingers
Ion pairs (salt bridges) stabilize local structure, while zinc fingers (Zn binding to His and Cys) are important for DNA binding.
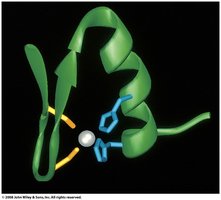
Protein Dynamics
Proteins are not static; they exhibit dynamic movements that are essential for function and catalysis. NMR studies reveal these dynamic properties.

Protein Denaturation and Renaturation
Denaturation
Proteins can be denatured by heat, pH changes, detergents, or chaotropic agents (e.g., urea, guanidinium ions). Denaturation disrupts weak bonding forces and leads to loss of structure and function.
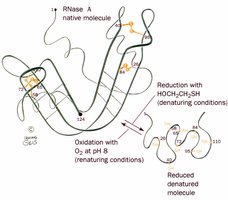
Renaturation
Some proteins, such as RNase A, can refold spontaneously after denaturation, demonstrating that primary structure dictates three-dimensional structure.
Protein Folding Pathways and Chaperones
Folding Pathways
Protein folding is not random; it follows directed pathways to reach the native conformation. The number of possible conformations is astronomically large, but proteins fold rapidly due to energy landscapes and assistance from chaperones.
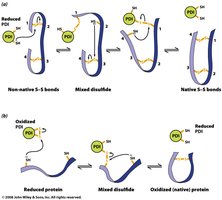
Disulfide Isomerase: Catalyzes rearrangement and formation of disulfide bonds.
Molecular Chaperones: Bind unfolded proteins and prevent improper folding.
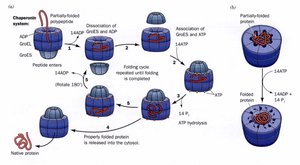
Protein Misfolding Diseases
Alzheimer's and Prion Diseases
Protein misfolding can lead to diseases such as Alzheimer's (misfolding of amyloid-β protein) and prion diseases (misfolding of PrP protein). Misfolded proteins aggregate into amyloid fibrils, causing cellular dysfunction and neurodegeneration.
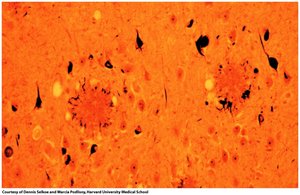
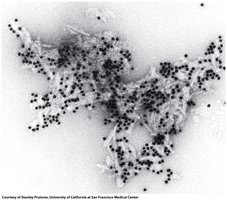
Transmissible Spongiform Encephalopathies (TSE)
TSEs are caused by prion proteins with abnormal folding. The misfolded form (PrPSC) induces misfolding in normal proteins (PrPC), leading to disease.