 Back
BackProteins: Primary Structure, Purification, and Sequencing
Study Guide - Smart Notes

Proteins: Primary Structure
The Sequence of Amino Acids in a Protein
The primary structure of a protein refers to the unique sequence of amino acids that make up the polypeptide chain. This sequence is encoded by the nucleotide sequence of DNA and is read from the amino (N) terminus to the carboxyl (C) terminus. The primary structure contains the information necessary for the protein to fold into its specific three-dimensional structure, which is essential for its function.
Genetic Information: The amino acid sequence is a direct translation of genetic information.
Directionality: Sequence is always written N-terminus to C-terminus.
Functional Implications: The primary structure determines protein folding and function.
Polypeptide Diversity
Although theoretically, the number of possible polypeptide sequences is enormous (20n for a chain of n residues), biological constraints limit this diversity. Protein synthesis efficiency and the requirement for proper folding restrict the size and composition of polypeptides.
Potential Variety: For a 100-residue protein, there are 1.27 x 10130 possible sequences.
Biological Limits: Most proteins are between 40 and 10,000 residues.
Fundamental Structural Pattern in Proteins
Proteins are unbranched polymers of amino acids joined head-to-tail by peptide bonds. Peptide bond formation is a condensation reaction, resulting in the release of water. The peptide backbone consists of a repeating sequence: –N–Cα–Co–.
Peptide Bond: Covalent bond formed between the carboxyl group of one amino acid and the amino group of another.
Residue: The term for an amino acid within a polypeptide chain, after water is eliminated.
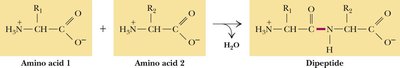
Peptide Bond Structure and Properties
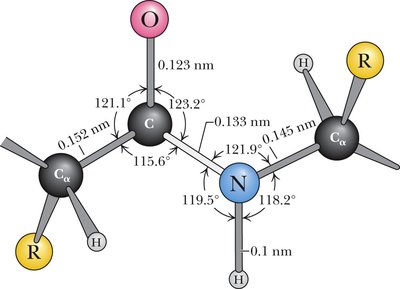
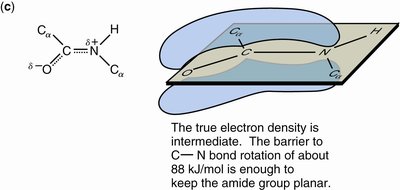
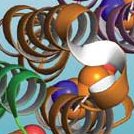
The peptide bond adopts a trans conformation and exhibits partial double bond character due to resonance. This restricts rotation and defines a planar structure known as the amide plane.
Bond Length: Approximately 0.133 nm, shorter than a single bond but longer than a double bond.
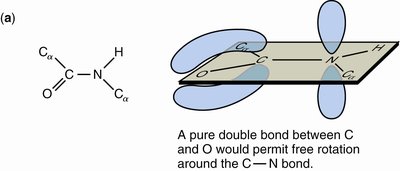
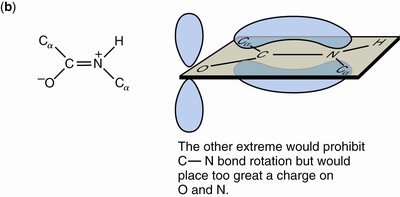
Resonance: The peptide bond is a resonance hybrid, contributing to its planarity and partial charges.
Hydrogen Bonding: N-H is a good donor; C=O is a good acceptor.
Peptide Nomenclature
Peptides are classified based on the number of amino acid residues:
Dipeptide: 2 residues
Tripeptide: 3 residues
Oligopeptide: 12–20 residues
Polypeptide: Many residues
Proteins may consist of one or more polypeptide chains:
Monomeric protein: Single polypeptide chain
Multimeric protein: Multiple chains (homomultimer: same type; heteromultimer: different types)
Example: Hemoglobin is a heterotetramer (2 alpha, 2 beta chains)
Protein Size and Composition
Proteins vary greatly in size, from small peptides to large multi-subunit complexes. Their molecular weights and isoelectric points (pI) are important for characterization.
Small Proteins: Insulin, Cytochrome c, Ribonuclease, Lysozyme, Myoglobin
Large Proteins: Hemoglobin, Immunoglobulin, Glutamine synthetase
Protein | Amino Acid Residues | Subunits | Protein Molecular Mass (D) |
|---|---|---|---|
Proteinase inhibitor III (bitter gourd) | 30 | 1 | 3,427 |
Cytochrome c (human) | 104 | 1 | 11,617 |
Myoglobin (horse) | 153 | 1 | 16,951 |
Interferon-γ (rabbit) | 288 | 2 | 33,482 |
Chorismate mutase (Bacillus subtilis) | 288 | 3 | 43,551 |
Triose phosphate isomerase (E. coli) | 510 | 2 | 53,944 |
Hemoglobin (human) | 574 | 4 | 64,764 |
RNA polymerase (bacteriophage T7) | 883 | 1 | 98,885 |
Nucleoside diphosphate kinase (Dictyostelium discoideum) | 2,252 | 4 | 245,456 |
Pyruvate decarboxylase (yeast) | 2,252 | 4 | 245,456 |
Glutamine synthetase (E. coli) | 5,616 | 12 | 621,264 |
Titin (mouse) | 35,213 | 1 | 3,906,488 |
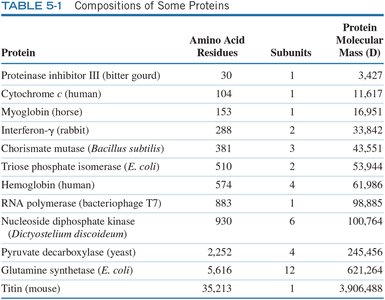
Protein Purification & Analysis
Environmental Conditions and Assays
Protein purification requires careful control of environmental conditions such as pH and temperature to maintain protein stability. Quantification is often achieved using assays based on chemical or binding properties, such as the enzyme-linked immunosorbent assay (ELISA).
Assay Example: ELISA uses antibodies to detect and quantify proteins.
Fractionation: Procedures exploit unique protein properties for separation.
Protein Purification Methods
Several methods are used to purify proteins based on their physical and chemical properties:
Salting Out: Separates proteins based on solubility in high salt concentrations.
Chromatography: Separates based on ionic charge, polarity, size, or ligand-binding ability.
Gel Electrophoresis: Separates proteins by charge, size, and isoelectric point.
Ultracentrifugation: Assesses size and shape of macromolecules.
Protein Characteristic | Purification Procedure |
|---|---|
Solubility | Salting out |
Ionic Charge | Ion exchange chromatography, Electrophoresis, Isoelectric focusing |
Polarity | Hydrophobic interaction chromatography |
Size | Gel filtration chromatography, SDS-PAGE |
Binding Specificity | Affinity chromatography |
Salting Out
Salting out is a technique where increasing salt concentration causes selective precipitation of proteins based on their solubility.
Principle: Proteins precipitate at different salt concentrations.
Application: Used to fractionate proteins from mixtures.
Ion Exchange Chromatography
Ion exchange chromatography separates proteins based on their net charge. Proteins bind to columns containing charged resins and are eluted by increasing salt concentration.
Cation Exchange: Positively charged proteins bind to negatively charged columns.
Anion Exchange: Negatively charged proteins bind to positively charged columns.
Dialysis
Dialysis is used to exchange buffer and remove small solutes such as salt from protein solutions. The sample is placed in a semipermeable membrane bag and immersed in a solution, allowing small molecules to equilibrate while retaining the protein.
Gel Filtration Chromatography
Gel filtration chromatography separates proteins based on size. Large proteins elute first because they cannot enter the pores of the gel beads, while small proteins are delayed.
Order of Elution: Largest to smallest
Affinity Chromatography
Affinity chromatography exploits specific binding interactions between a protein and a ligand immobilized on a column. Only proteins with affinity for the ligand are retained and can be eluted by adding excess ligand.
Electrophoresis and SDS-PAGE
Electrophoresis separates proteins by charge and size. SDS-PAGE uses sodium dodecyl sulfate to denature proteins and impart a uniform negative charge, allowing separation by molecular weight.
SDS-PAGE: Mobility is inversely proportional to molecular weight.
2-D Gel Electrophoresis: Separates proteins first by mass, then by isoelectric point.
Protein Sequencing
Overview of Protein Sequencing
Protein sequencing involves determining the order of amino acids in a polypeptide. The process requires separation of chains, cleavage of disulfide bonds, identification of terminal residues, fragmentation, and reconstruction of the sequence.
Edman Degradation: Sequential removal of N-terminal residues for identification.
Mass Spectrometry: Identifies sequences based on mass-to-charge ratios.
Databases: Sequence data are stored in online databases for interpretation.
Steps in Protein Sequencing
Separate polypeptide chains by denaturation.
Cleave disulfide bonds using reducing agents (e.g., β-mercaptoethanol, DTT) and alkylate to prevent reformation.
Identify N-terminal (Edman's reagent) and C-terminal residues (carboxypeptidases).
Fragment chains using enzymatic (trypsin, chymotrypsin, staphylococcal protease) or chemical (cyanogen bromide) methods.
Repeat fragmentation with different agents to generate overlapping fragments.
Reconstruct the sequence from overlapping fragments.
Enzymatic and Chemical Fragmentation
Trypsin: Cleaves C-side of Lys and Arg.
Chymotrypsin: Cleaves C-side of Phe, Tyr, Trp.
Staphylococcal Protease: Cleaves C-side of Glu, Asp.
Cyanogen Bromide: Cleaves at Met residues, producing peptides with C-terminal homoserine lactone.
Mass Spectrometry
Mass spectrometry separates protein fragments based on mass-to-charge ratio, allowing identification of peptide sequences. ESI-MS and MALDI-TOF MS are commonly used techniques.
Peptide Mass Fingerprinting: Matches fragment masses to database entries.
Protein Sequence Databases
Protein and DNA sequence data are stored in online databases, facilitating identification and comparison of protein sequences.
Summary Table: Protein Sequencing Toolbox
Reagent/Enzyme | Function |
|---|---|
β-mercaptoethanol (β-Me), DTT | Reduces disulfide bonds |
Iodoacetate | Alkylates thiols to prevent disulfide reformation |
Edman's reagent | N-terminal sequencing |
Carboxypeptidase A/B | C-terminal sequencing |
Trypsin | Cleaves C-side of Lys, Arg |
Chymotrypsin | Cleaves C-side of Phe, Tyr, Trp |
Staphylococcal protease | Cleaves C-side of Glu, Asp |
Cyanogen bromide (CNBr) | Cleaves at Met residues |
Additional info: The notes have been expanded with academic context to ensure completeness and clarity for biochemistry students.
