 Back
BackProteins: Three-Dimensional Structure – Study Notes
Study Guide - Smart Notes

Proteins: Three-Dimensional Structure
Overview of Protein Structure
Proteins are complex macromolecules essential for life, and their function is determined by their three-dimensional structure. Protein structure is organized hierarchically into four levels: primary, secondary, tertiary, and quaternary. Each level of structure is stabilized by specific types of chemical interactions.
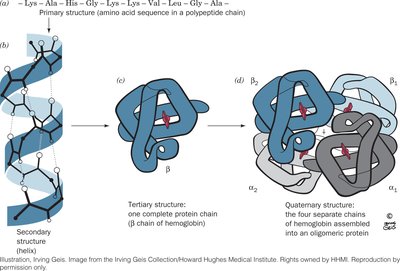
Primary Structure (1°): The linear sequence of amino acids in a polypeptide chain, held together by covalent peptide bonds.
Secondary Structure (2°): Local folding patterns such as α helices and β sheets, stabilized by hydrogen bonds between backbone atoms.
Tertiary Structure (3°): The overall three-dimensional shape of a single polypeptide, stabilized by various weak interactions.
Quaternary Structure (4°): The arrangement of multiple polypeptide subunits in a protein complex.
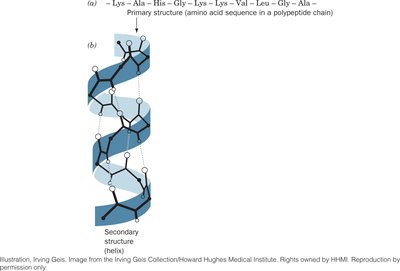
Hierarchy of Protein Structure
Globular Proteins: Spherical, soluble proteins with diverse functions.
Fibrous Proteins: Elongated, insoluble proteins with structural roles.
The four levels of protein structure are illustrated below, using hemoglobin as an example:
Stabilizing Forces in Protein Structure
Secondary, tertiary, and quaternary structures are stabilized by weak, noncovalent interactions:
Hydrogen Bonds: Formed wherever possible, directional and crucial for secondary structure.
Hydrophobic Interactions: Drive protein folding by increasing water entropy.
Ionic Interactions: Abundant on protein surfaces, occur between oppositely charged groups.
Van der Waals Interactions: Ubiquitous, optimize packing within the protein.
Secondary Structure
Constraints of Peptide Chemistry
The peptide bond is planar due to resonance stabilization, restricting rotation. However, rotation is allowed around the bonds adjacent to the α-carbon, defined by the torsion angles φ (phi) and ψ (psi).
φ (phi): Rotation around the N–Cα bond.
ψ (psi): Rotation around the Cα–C bond.
Not all φ and ψ combinations are allowed due to steric hindrance. The Ramachandran plot shows the sterically allowed regions for these angles, corresponding to common secondary structures.
Types of Secondary Structure
α Helix: Right-handed helix stabilized by hydrogen bonds between the carbonyl oxygen of residue i and the amide hydrogen of residue i+4. Typical values: 3.6 residues per turn, 1.5 Å rise per residue, pitch of 5.4 Å.
β Sheet: Composed of β strands connected laterally by at least two or three backbone hydrogen bonds, forming a sheet-like structure. Strands can be parallel or antiparallel.
β Turn: Allows the polypeptide chain to reverse direction, stabilized by a hydrogen bond between the carbonyl oxygen of residue i and the amide hydrogen of residue i+3.
Helix-Forming and Helix-Breaking Amino Acids
Certain amino acids favor or disrupt α helix formation. For example, proline is a helix breaker due to its rigid structure, while glycine is too flexible.
Amino Acid | Helix Behavior |
|---|---|
Ala | H (I) |
Pro | B |
Gly | B |
Leu | H (I) |
Ser | H (D) |
Thr | Variable |
Fibrous Proteins
α Keratin
Found in hair, nails, and other structures, α keratin consists of coiled coils with a heptad repeat (a-b-c-d-e-f-g)n, where a and d are nonpolar residues. This structure promotes the association of α helices into stable fibers.
β Keratin
Present in silk and feathers, β keratin forms stacked β sheets with an alternating sequence (Gly-Ala/Ser)n. The close packing of glycine and alanine/serine residues provides strength through van der Waals interactions.
Collagen
Collagen is the main component of connective tissue. It forms a triple helix with a repeating (Gly-Pro-HyPro)n sequence. The structure is stabilized by interchain hydrogen bonds and covalent crosslinks.
Tertiary Structure
Principles of Tertiary Structure
Tertiary structure refers to the overall three-dimensional arrangement of all atoms in a protein. It is determined by the folding and packing of secondary structural elements into motifs and domains. Nonpolar residues are typically buried in the interior, while polar residues are exposed to the solvent.
Motifs: Small folding patterns found in many proteins.
Domains: Independently stable and functional regions within a protein.
Protein Surface and Core
The core is mainly hydrophobic and contains α helices and β sheets.
The surface is more variable, with loops and turns that interact with other molecules.
Water molecules are often found on the protein surface, forming hydrogen bonds with polar groups.
Quaternary Structure
Subunit Organization and Symmetry
Proteins with quaternary structure contain multiple polypeptide chains (subunits) arranged symmetrically. Subunit interactions are stabilized by the same weak forces as tertiary structure. Quaternary structure provides stability, genetic economy, assembly of catalytic sites, and cooperativity.
Protein Stability and Folding
Stability
Protein stability is primarily due to hydrophobic effects, with additional contributions from hydrogen bonds and electrostatic interactions. Proteins are only marginally stable, with a typical folding free energy (ΔG) of -5 to -10 kJ/mol, allowing for necessary flexibility and motion.
Denaturation and Renaturation
Denaturation is the loss of protein structure and function due to external stresses such as heat or chemicals. Some proteins can refold (renature) to regain their native structure and function, as demonstrated by ribonuclease A.
Protein Folding Pathways
Protein folding is a complex process that does not occur by random sampling of all possible conformations (Levinthal's paradox). Instead, folding follows specific pathways, often involving intermediate states such as the molten globule. Molecular chaperones assist in proper folding and prevent aggregation.
Folding Funnel Model: The folding process can be visualized as a funnel of decreasing free energy and conformational possibilities.
Chaperones: Proteins that assist in folding, especially under stress conditions.
Summary Table: Key Features of Protein Structure
Level | Definition | Stabilizing Forces |
|---|---|---|
Primary | Amino acid sequence | Peptide bonds (covalent) |
Secondary | Local folding (α helix, β sheet) | Hydrogen bonds |
Tertiary | 3D arrangement of polypeptide | Hydrophobic, H bonds, ionic, van der Waals |
Quaternary | Assembly of multiple subunits | Same as tertiary |
Additional info: For a deeper understanding, students should be able to interpret Ramachandran plots, identify motifs and domains, and explain the significance of protein folding diseases and the role of chaperones.
