 Back
BackRegulation of Gene Expression: Prokaryotic and Eukaryotic Mechanisms
Study Guide - Smart Notes

Regulation of Gene Expression
Overview
The regulation of gene expression is a fundamental process that allows cells to control the production of proteins and enzymes in response to internal and external signals. This regulation is essential for cellular efficiency, adaptation, and differentiation. In both prokaryotes and eukaryotes, gene expression is tightly controlled at multiple levels, including transcription, translation, and post-translational modifications.
Division of Genes in Living Organisms
Constitutive vs. Expressible Genes
Constitutive (Housekeeping) Genes: Genes that are expressed continuously to maintain basic cellular function.
Expressible Genes: Inducible or repressible genes that are only expressed when needed, allowing the cell to conserve energy and resources.
Examples of expressible genes include those involved in tanning, muscle protein synthesis in response to exercise, and genes activated during pregnancy.
Prokaryotic Gene Regulation: The lac Operon
Operons and Coordinated Gene Expression
In prokaryotes, genes involved in related functions are often organized into operons, which are clusters of genes regulated together by a single promoter. This arrangement allows for coordinated expression in response to environmental changes.
The Lactose (lac) Operon in E. coli
The lac operon is a classic example of gene regulation in prokaryotes. It enables E. coli to metabolize lactose only when it is available, thereby conserving energy when lactose is absent.
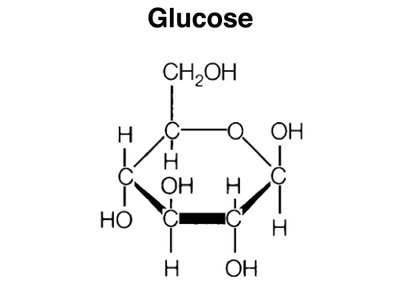
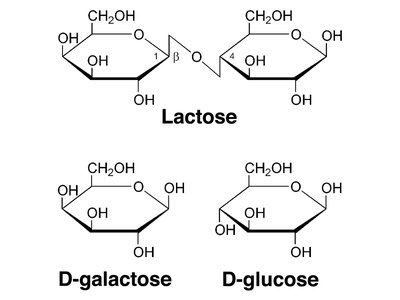
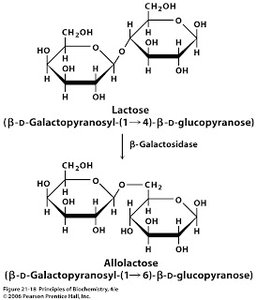
Lactose: A disaccharide composed of glucose and galactose.
lac Operon Genes: lacZ (β-galactosidase), lacY (permease), lacA (transacetylase).
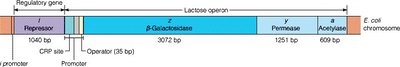
Organization of the lac Operon
The lac operon consists of a promoter, operator, and three structural genes. The promoter is the binding site for RNA polymerase, while the operator is the binding site for the lac repressor protein.
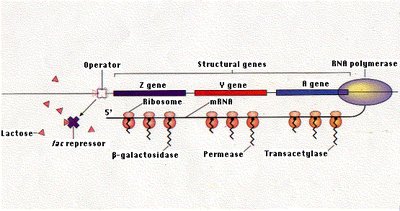
Regulation by the lac Repressor
No Lactose Present: The lac repressor binds to the operator, blocking RNA polymerase and preventing transcription (repression).
Lactose Present: An isomer of lactose, allolactose, binds to the repressor, causing it to dissociate from the operator. This allows transcription of the operon and synthesis of enzymes needed for lactose metabolism.
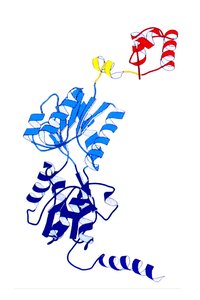
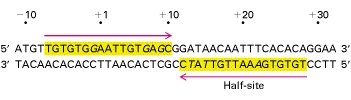
lac Repressor Protein and Operator Sequence
The lac repressor is a 360-amino acid protein that binds specifically to a 35 base pair palindromic sequence in the operator region, preventing transcription.

Catabolite Activator Protein (CAP) and cAMP
Transcription of the lac operon is also regulated by the availability of glucose. When glucose is scarce, cyclic AMP (cAMP) levels rise, allowing cAMP to bind to CAP. The CAP-cAMP complex binds upstream of the promoter, enhancing RNA polymerase binding and transcription. If glucose is present, cAMP levels are low, CAP does not bind, and transcription is minimal even if lactose is present.
Regulation of Transcription in Eukaryotes
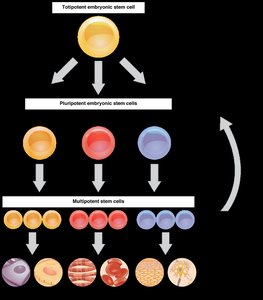
Spatial and Temporal Expression
In multicellular organisms, gene expression is regulated so that different cell types express different sets of genes (spatial expression), and genes are expressed at appropriate times (temporal expression). Misregulation can lead to disease.
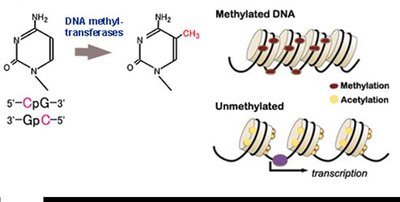
Mechanisms of Eukaryotic Gene Regulation
DNA Methylation: Addition of methyl groups to cytosine bases in DNA, leading to gene silencing by preventing transcription factor binding.
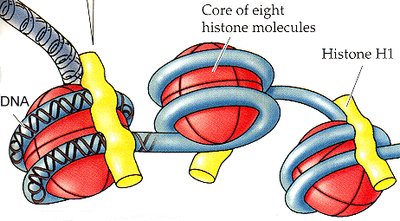
Histone Acetylation: Addition of acetyl groups to histone proteins, causing chromatin to unwind and making DNA more accessible for transcription.
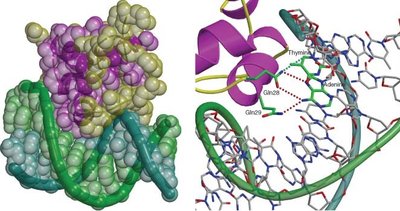
Transcription Factors
Transcription factors are proteins that bind to specific DNA sequences, facilitating or inhibiting the recruitment of RNA polymerase and other components of the transcriptional machinery. They play a central role in determining which genes are expressed in a given cell type.
Summary Table: Key Features of Prokaryotic and Eukaryotic Gene Regulation
Feature | Prokaryotes (lac Operon) | Eukaryotes |
|---|---|---|
Gene Organization | Operons (clusters of genes under one promoter) | Genes usually regulated individually |
Regulatory Proteins | Repressors, activators (e.g., CAP) | Transcription factors, co-activators, repressors |
Regulatory Elements | Promoter, operator, CAP site | Promoters, enhancers, silencers |
Epigenetic Regulation | Rare | Common (DNA methylation, histone modification) |
Response to Environment | Rapid, direct (e.g., presence of lactose or glucose) | Complex, involves multiple layers of regulation |
