 Back
BackThe Three-Dimensional Structure of Proteins: Secondary, Tertiary, and Quaternary Organization
Study Guide - Smart Notes

Three-Dimensional Structure of Proteins
Overview of Protein Structure
Proteins exhibit a hierarchical structure, classified into four distinct levels: primary, secondary, tertiary, and quaternary. Each level contributes to the overall function and stability of the protein molecule.
Primary Structure: The linear sequence of amino acids in a polypeptide chain.
Secondary Structure: Localized, repeating structures stabilized by hydrogen bonds, such as α-helices and β-sheets.
Tertiary Structure: The overall three-dimensional folding of a single polypeptide chain, resulting from interactions among secondary structural elements.
Quaternary Structure: The spatial arrangement of multiple polypeptide chains (subunits) in a multi-subunit complex.
Pauling’s Rules for Secondary Structure
Linus Pauling and collaborators established foundational principles for secondary protein structure:
Bond angles and lengths should closely resemble those of free amino acids.
No two atoms should approach closer than their van der Waals radii allow (steric restrictions).
The six atoms in the peptide amide group must remain coplanar with associated α-carbons in the trans configuration.
Non-covalent bonding, especially hydrogen bonding, is essential for stabilizing regular folding patterns.
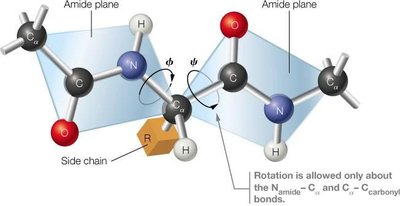
Secondary Structure: α-Helices and β-Sheets
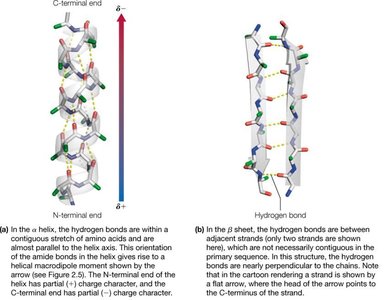
Secondary structures are stabilized by hydrogen bonds between backbone amide protons and carbonyl oxygens. The two most prevalent secondary structures are the right-handed α-helix and the β-sheet.
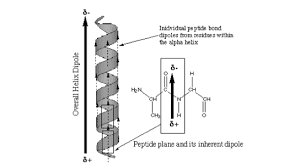
α-Helix: A helical structure with 3.6 amino acid residues per turn. Side chains project outward from the helix.
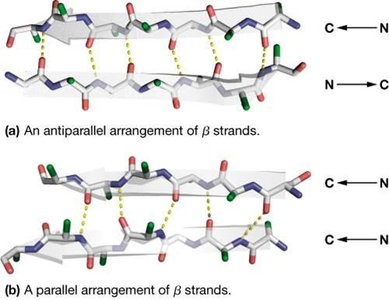
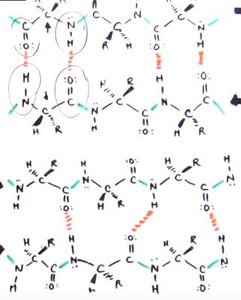
β-Sheet: Composed of two or more β-strands, stabilized by interchain hydrogen bonds. Side chains alternate above and below the sheet.
β-strands can be arranged in parallel (both strands N→C) or antiparallel (one strand N→C, the other C→N) configurations.
Hydrogen bonding patterns differ: antiparallel sheets have 1:1 H-bonding, while parallel sheets have more complex bonding.
Polyproline II Helix
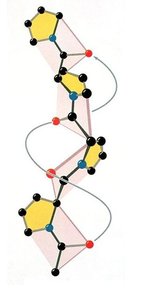
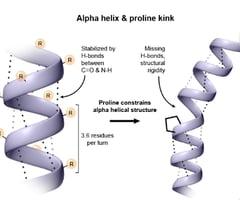
The polyproline II helix is a left-handed helix commonly found in protein structures. It does not satisfy Pauling’s requirements for hydrogen bonding and is stabilized primarily by steric factors. Proline and glycine residues are frequent, with proline causing kinks and glycine providing flexibility.
Amphipathic Helices and Sheets
Many α-helices and β-sheets are amphipathic, meaning they have distinct hydrophobic and hydrophilic faces. This property allows secondary structures to associate via hydrophobic interactions while exposing hydrophilic surfaces to the aqueous environment.
Fibrous Proteins: Structural Materials of Cells and Tissues
Characteristics and Types of Fibrous Proteins
Fibrous proteins are elongated molecules with well-defined secondary structures, serving structural roles in cells and tissues. They are enriched in specific amino acids that stabilize their extended conformations.
Keratin: Major protein in hair, fingernails, feathers, and skin. Predominantly α-helical.
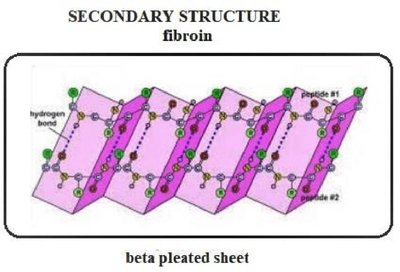
Fibroin: Protein in silk, composed mainly of β-sheets.
Collagen: Abundant connective tissue protein, forms the matrix in bone and tendons.
Amino Acid Composition of Fibrous Proteins
Fibrous proteins have a high abundance of certain amino acids, which confer mechanical properties and stability.
Amino Acid | α-Keratin (wool) | Fibroin (silk) | Collagen (bovine tendon) | All Proteins |
|---|---|---|---|---|
Gly | 8.1 | 44.6 | 32.7 | 7.9 |
Ala | 5.0 | 29.4 | 12.0 | 8.7 |
Ser | 10.2 | 12.2 | 3.4 | 5.8 |
Glu + Gln | 12.1 | 1.0 | 7.7 | 6.6 |
Cys | 11.2 | 0 | 0 | 1.3 |
Pro | 7.5 | 0.3 | 12.1 | 4.7 |
Arg | 7.2 | 0.5 | 5.0 | 5.0 |
Leu | 6.9 | 0.5 | 2.1 | 8.9 |
Thr | 6.5 | 0.9 | 1.6 | 5.6 |
Asp + Asn | 6.0 | 1.3 | 4.5 | 5.9 |
Val | 5.1 | 2.2 | 1.8 | 7.2 |
Tyr | 4.2 | 5.2 | 0.4 | 3.5 |
Ile | 2.8 | 0.7 | 0.9 | 5.5 |
Phe | 2.5 | 0.5 | 1.2 | 4.0 |
Lys | 2.3 | 0.3 | 3.7 | 5.5 |
Trp | 1.2 | 0.2 | 0 | 1.5 |
His | 0.7 | 0.2 | 0.3 | 2.4 |
Met | 0.5 | 0 | 0.7 | 2.0 |
Keratins
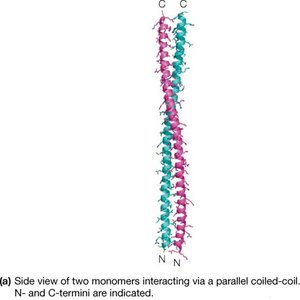
α-Keratins are major proteins of hair and fingernails, consisting of long α-helical sequences. Pairs of right-handed helices twist about one another in a left-handed coiled-coil structure, stabilized by hydrophobic interactions and disulfide cross-links.
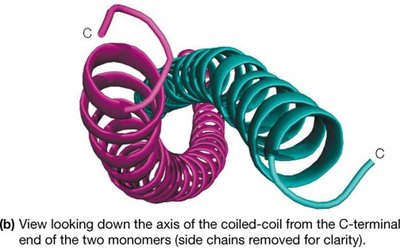
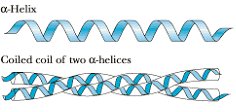
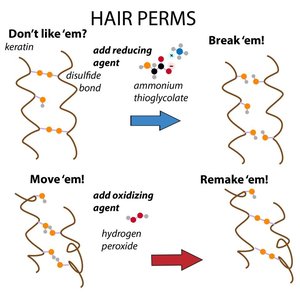
Fibroin
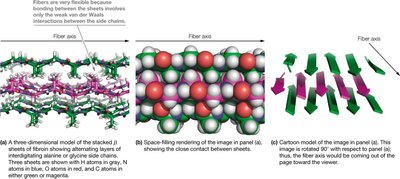
Fibroin, the protein in silk, utilizes stacked antiparallel β-sheets. The sheets are held together by non-covalent interactions between interdigitated side chains, primarily glycine, alanine, and serine. This structure provides strength and flexibility.
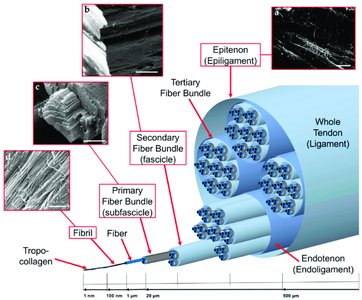
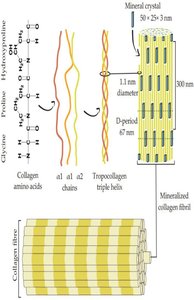
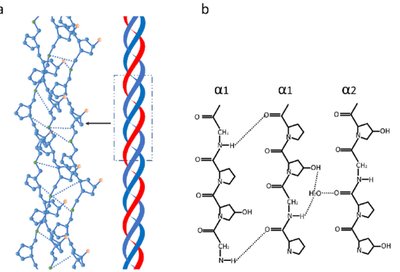
Collagen
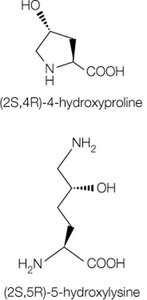
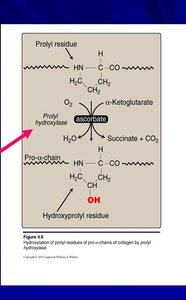
Collagen is the most abundant protein in vertebrates, forming the matrix in bone, tendons, and skin. The basic unit, tropocollagen, is a triple helix of three polypeptide chains, each about 1000 residues long. The triple helix is stabilized by hydrogen bonds and modifications such as hydroxylation of proline and lysine.

Globular Proteins: Tertiary Structure and Functional Diversity
Principles of Protein Folding
Globular proteins fold into compact, three-dimensional structures. Folding is driven by the formation of secondary structures, packing of helices and sheets, and clustering of hydrophobic groups in the protein interior.
Secondary structures form wherever possible due to hydrogen bonding.
Helices and sheets pack closely together.
Peptide segments between secondary structures are short and direct.
Stability arises from intramolecular hydrogen bonds and reduction in solvent-accessible surface area.
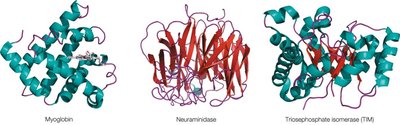
Varieties of Globular Protein Structure
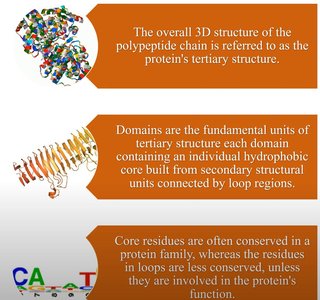
Globular proteins exhibit diverse folding patterns, often classified by dominant secondary structure motifs. Domains are compact, locally folded regions of tertiary structure, often performing distinct functions.

Factors Determining Secondary and Tertiary Structure
Information for Protein Folding
The amino acid sequence contains most of the information required for protein folding. Denaturation, the loss of structure and function, can be caused by changes in pH, temperature, or the addition of organic solvents.
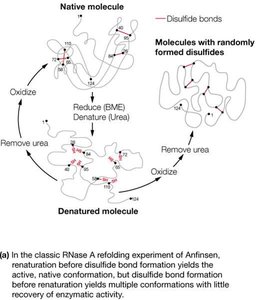
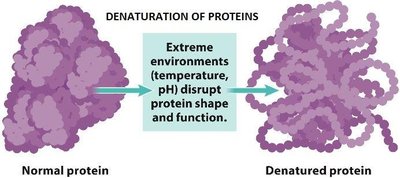
Thermodynamics, Folding, and Stability of Proteins
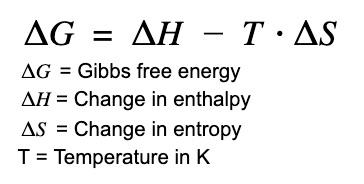
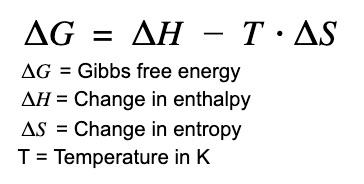
Protein folding is a thermodynamically favorable process under physiological conditions. The free energy change () for folding is determined by enthalpy () and entropy ():
Conformational Entropy: Folding reduces randomness (entropy), which opposes folding.
Charge-Charge Interactions: Electrostatic attractions and repulsions affect stability.
Internal Hydrogen Bonds: Numerous hydrogen bonds among side chains and backbone stabilize the folded state.
van der Waals Interactions: Weak interactions among densely packed non-polar groups contribute to stability.
Hydrophobic Effect: Burial of hydrophobic side chains increases solvent entropy, favoring folding.


Binding of Ions or Prosthetic Groups
Protein stability can be increased by binding metal ions or cofactors. The active, stable form is called a holoprotein, while the inactive form lacking the cofactor is an apoprotein.
Quaternary Structure of Proteins
Oligomeric Proteins and Subunit Organization
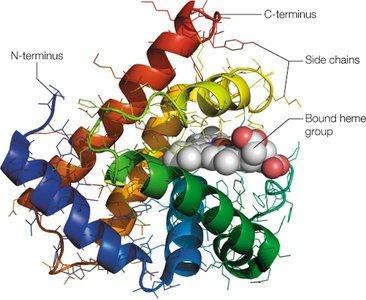
Proteins with two or more polypeptide chains are termed oligomers and possess quaternary structure. Subunits are held together by noncovalent bonds. Examples include hemoglobin, aspartate transcarbamylase, and lactate dehydrogenase.
Hemoglobin: A tetrameric protein in red blood cells, consisting of two α and two β chains, each with a heme group capable of binding oxygen.
Models of Protein Folding and Aggregation
Protein Misfolding and Disease
Misfolding of proteins can lead to aggregation and disease. Amyloid fibrils are highly ordered aggregates characterized by β-sheet structures. Prion diseases are caused by infectious protein aggregates that induce amyloid formation.
Amyloid Fibrils: Formed from non-native folding intermediates or disordered aggregate states.
Prions: Infectious agents that cause disease by inducing amyloid formation upon contact.
Disease | Associated Protein |
|---|---|
Alzheimer’s disease | Amyloid β peptide |
Parkinson’s disease | α-Synuclein |
Spongiform encephalopathies (CJD, kuru) | Prion protein |
Amyotrophic lateral sclerosis (ALS) | Superoxide dismutase I |
Huntington’s disease | Huntingtin with polyQ tracts |
Cataracts | γ-Crystallin |
Type II diabetes | Islet amyloid polypeptide (IAPP) |
Injection-localized amyloidosis | Insulin |
Summary
The three-dimensional structure of proteins is essential for their function and stability. Understanding the principles of protein folding, the roles of secondary, tertiary, and quaternary structures, and the consequences of misfolding is fundamental in biochemistry.
