 Back
BackThree-Dimensional Structure of Proteins: Secondary and Tertiary Structure
Study Guide - Smart Notes

Three-Dimensional Structure of Proteins
Secondary Structure: Overview
Secondary structure refers to the local spatial arrangement of the polypeptide backbone, stabilized primarily by hydrogen bonds. The most common secondary structures are the α-helix and β-sheet, with turns and loops connecting these elements. Understanding secondary structure is essential for predicting protein folding and function.
Helical Propensity of Amino Acids
The likelihood of an amino acid to be found in an α-helix, β-sheet, or turn is termed its secondary structure propensity. This is influenced by the side chain's size, charge, and branching.
Amino Acid | α helix | β sheet | Turn |
|---|---|---|---|
Ala | 1.29 | 0.90 | 0.78 |
Cys | 1.11 | 1.19 | 0.74 |
Leu | 1.34 | 1.10 | 0.66 |
Met | 1.44 | 1.05 | 0.60 |
Glu | 1.22 | 0.75 | 1.01 |
His | 1.24 | 0.87 | 0.87 |
Lys | 1.23 | 0.74 | 1.01 |
Phe | 1.20 | 1.29 | 0.60 |
Gln | 1.27 | 0.80 | 0.97 |
Val | 0.88 | 1.65 | 0.57 |
Ile | 0.97 | 1.60 | 0.47 |
Tyr | 0.77 | 1.38 | 0.69 |
Trp | 1.08 | 1.37 | 0.54 |
Thr | 0.82 | 1.19 | 1.20 |
Ser | 0.77 | 0.72 | 1.32 |
Asp | 0.54 | 0.70 | 1.33 |
Asn | 0.76 | 0.65 | 1.56 |
Arg | 1.21 | 0.92 | 0.93 |
Pro | 0.57 | 0.63 | 1.32 |
Gly | 0.53 | 0.64 | 1.54 |

Example: Alanine is highly favored in α-helices, while Glycine and Proline are more common in turns due to their unique backbone flexibility.
Helical Wheel Views and Heptad Repeats
A helical wheel is a graphical representation of the spatial arrangement of amino acids in an α-helix, viewed down the helical axis. This helps visualize amphipathic helices, where one side is hydrophobic and the other is hydrophilic.
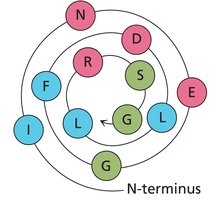
Heptad repeats are patterns of seven amino acids (a-g) that repeat, often found in coiled-coil structures. Positions a and d are typically hydrophobic, facilitating helix-helix interactions.
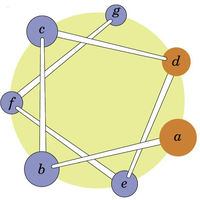
Example: Drawing a helical wheel for the peptide segment HELICAL can reveal which residues are likely to interact with the environment or other helices.
Electrostatic Interactions in Helices
Electrostatic interactions between side chains at positions i and i+3 or i and i+4 stabilize the helix. These interactions can be ionic or hydrophobic, depending on the nature of the side chains.
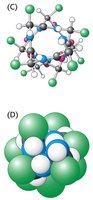
Example: Lysine (positive) and Glutamate (negative) at i and i+4 positions can form a stabilizing salt bridge.
Supersecondary Structure: Coiled-Coil
Coiled-coil motifs are formed when two or more α-helices wind around each other, stabilized by hydrophobic interactions at specific positions (usually a and d in the heptad repeat). These structures are common in fibrous proteins and play roles in structural integrity and protein-protein interactions.
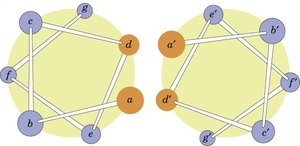
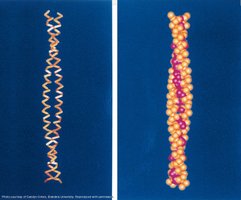
Example: The coiled-coil motif is found in proteins such as myosin and tropomyosin.
Sequence Rules for α-Helix Formation
Several factors influence α-helix formation:
Helical propensity: Linear, non-β-branched amino acids are favored; Proline and Glycine are rare due to steric constraints.
Adjacent interactions: Bulky side chains near the backbone can hinder helix formation.
Electrostatics: Opposite charges may cluster, while similar charges repel.
Down the helical wheel: i to i+3 positions are closest and often interact via ionic or hydrophobic forces.
Nonstandard Helices
Helices can vary in their hydrogen bonding patterns, resulting in different numbers of residues per turn (e.g., 310 helix, π-helix). These are less common than the α-helix.
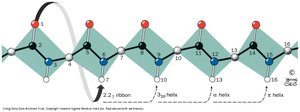
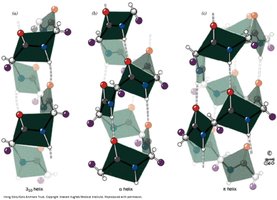
Example: The 310 helix has three residues per turn and tighter hydrogen bonding compared to the α-helix.
β-Sheet Structure
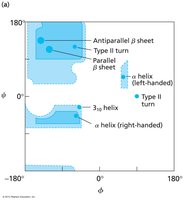
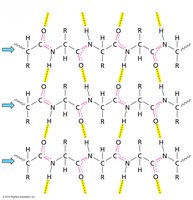
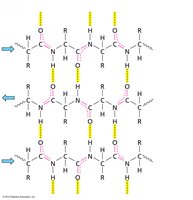
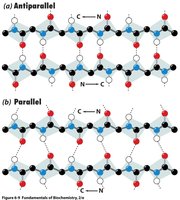
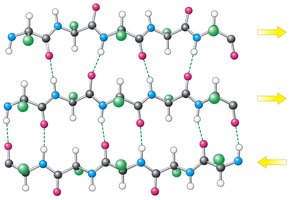
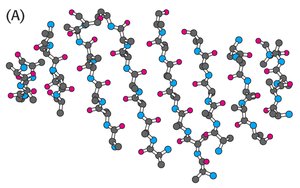
β-sheets are formed by extended polypeptide chains aligned side-by-side, stabilized by hydrogen bonds between backbone atoms. Sheets can be parallel or antiparallel, depending on the directionality of the strands.
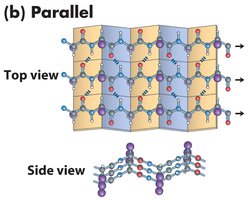
Parallel β-sheet: Strands run in the same direction; hydrogen bonds are non-linear.
Antiparallel β-sheet: Strands run in opposite directions; hydrogen bonds are linear and stronger.
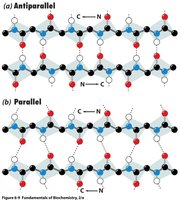
Example: Silk fibroin is composed of antiparallel β-sheets, giving it strength and flexibility.
β-Sheet Side Chain Arrangement
In β-sheets, side chains alternate above and below the plane of the sheet, minimizing steric clash and allowing for large, branched side chains to be accommodated.
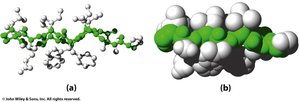
Example: Valine and Isoleucine are common in β-sheets due to their branched side chains.
β-Sheet Propensity of Amino Acids
Some amino acids are more likely to be found in β-sheets due to their side chain properties.
Example: Isoleucine and Valine have high β-sheet propensities.
Mixed and Extended β-Sheets
β-sheets can be mixed (containing both parallel and antiparallel strands) and are often twisted in three dimensions. Arrows indicate the N to C directionality of the strands.
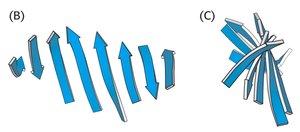
Example: The β-barrel motif in proteins such as GFP is formed by twisted β-sheets.
Irregular Structure: Turns and Loops
Turns and loops are nonstandard secondary structures that connect α-helices and β-sheets. Turns are short, often stabilized by hydrogen bonds, while loops are longer and more flexible.
Random coil: No detectable structure, high flexibility.
Coil: Structured regions not fitting standard definitions.
β-hairpin turns: Connect antiparallel β-strands, stabilized by hydrogen bonds between the first and fourth residues.
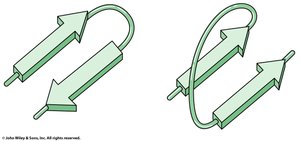
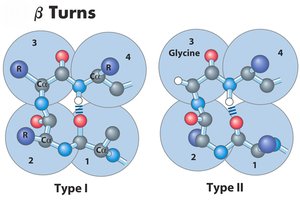

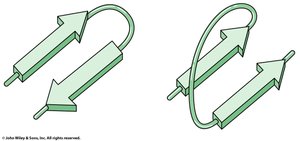
Example: Glycine and Proline are common in turn regions; Proline often appears in Type I turns, Glycine in Type II.
Summary of Secondary Structure
Proteins typically contain approximately one-third α-helices, one-third β-sheets, and one-third irregular structure. However, this ratio varies for individual proteins.
Tertiary Structure and Protein Folding
Common Depictions of Protein Structure
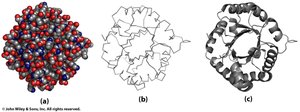
Protein structures can be visualized in several ways:
Space-filling (ball): Shows the volume occupied by atoms.
Backbone wire: Highlights the polypeptide backbone.
Backbone secondary structure: Emphasizes α-helices and β-sheets.
Protein Structure and Stability
Proteins have a hydrophobic core and a hydrophilic surface. Hydrophobic amino acids are buried inside, while charged and polar residues are exposed to the solvent.
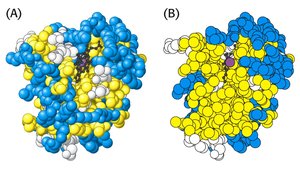
Example: Myoglobin's interior is hydrophobic, stabilizing its folded structure.
RNase A Structure and Anfinsen's Experiments
Christian Anfinsen demonstrated that the primary sequence of a protein contains all the information required for folding into its native, functional structure. His experiments with RNase A showed that proteins can spontaneously refold after denaturation if the correct conditions are provided.
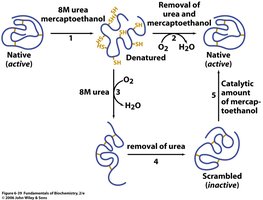
Example: RNase A, with four disulfide bonds, can regain activity after denaturation and renaturation.
Protein Folding Pathways
Protein folding occurs in two phases:
Fast phase: Local structural nucleation and hydrophobic collapse to a molten globule state.
Slow phase: Stabilization of secondary structure, packing of side chains, and expulsion of water from the core.
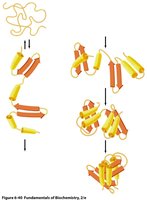
Folding funnel: Proteins fold via multiple pathways, reducing free energy and entropy until the native state is reached.
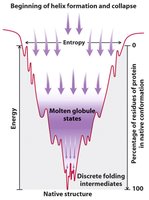
Driving Forces for Protein Folding
The hydrophobic effect is the primary driving force for protein folding, driven by entropy. Hydrogen bonds help define structure but are not the main source of stability due to desolvation costs.
Ion pairs and salt bridges: Interior salt bridges contribute specificity and stability, while exterior salt bridges are less significant due to water competition.
Protein Denaturation
Denaturation is the loss of secondary and tertiary structure, resulting in a random coil. The melting temperature (Tm) is the point at which this transition occurs. Denaturation can be caused by heat or chemical agents (chaotropes).
Example: Albumin denaturation is monitored by circular dichroism, showing a cooperative two-state transition.
Protein Misfolding and Disease
Misfolded proteins can aggregate and cause diseases such as Alzheimer's, Huntington's, and prion diseases. Prion proteins can catalyze the conversion of normal proteins to the misfolded, infectious form.
Molecular Chaperones
Molecular chaperones assist in proper protein folding by preventing aggregation and providing a framework for folding. Chaperonins such as GroEL/GroES use ATP hydrolysis to promote correct folding.
Example: GroEL/GroES complex is a classic chaperonin system studied in protein folding.
Key Concepts for Chapter 6
Understand why rotation of protein backbones is limited (steric hindrance, peptide bond planarity).
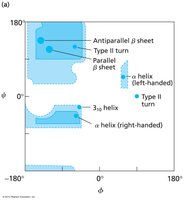
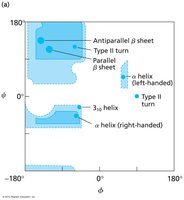
Interpret Ramachandran plots for allowed backbone conformations.
Use heptad repeats to represent helices and predict their properties.
Draw parallel and antiparallel β-strands.
Identify Type I and Type II turns in protein structures.
