 Back
BackTranscription and RNA Processing: Biochemistry Study Notes
Study Guide - Smart Notes

Transcription and RNA Processing
Introduction to Genes and RNA
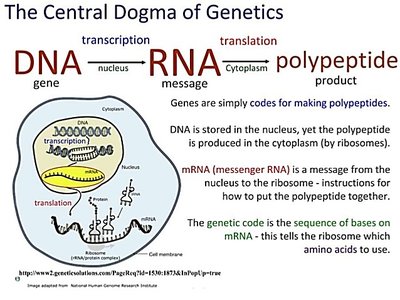
Genes are segments of DNA that are transcribed into RNA, serving as the fundamental units of heredity. The process of transcription converts genetic information from DNA into RNA, which can then be translated into proteins or function directly in the cell. There are several types of RNA, each with distinct roles in gene expression and regulation.
Gene: A DNA segment that is transcribed into RNA. Genes may encode proteins, transfer RNA (tRNA), or ribosomal RNA (rRNA).
Central Dogma: The flow of genetic information from DNA to RNA (transcription) and from RNA to protein (translation).
Types of RNA and Their Functions
RNA molecules play diverse roles in the cell, with three primary types involved in gene expression:
Messenger RNA (mRNA): Encodes the amino acid sequence of proteins; short-lived and constitutes a small fraction of total RNA.
Transfer RNA (tRNA): Delivers amino acids to ribosomes during protein synthesis; highly stable.
Ribosomal RNA (rRNA): Structural and catalytic component of ribosomes; highly stable.
Transcription: Mechanism and Regulation
Overview of Transcription
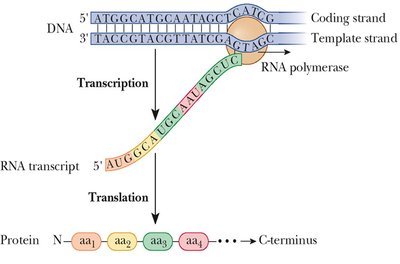
Transcription is the process by which RNA is synthesized from a DNA template. It involves three main stages: initiation, elongation, and termination. The enzyme RNA polymerase catalyzes the synthesis of RNA in the 5' to 3' direction, using one strand of DNA as a template.
Comparison: Replication vs. Transcription
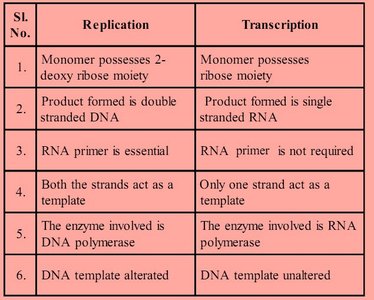
While both replication and transcription involve copying genetic information, they differ in several key aspects:
Replication | Transcription |
|---|---|
Monomer possesses 2-deoxy ribose moiety | Monomer possesses ribose moiety |
Product formed is double-stranded DNA | Product formed is single-stranded RNA |
RNA primer is essential | RNA primer is not required |
Both strands act as templates | Only one strand acts as a template |
Enzyme: DNA polymerase | Enzyme: RNA polymerase |
DNA template altered | DNA template unaltered |
Enzymology of Transcription
RNA polymerase is the key enzyme in transcription. It synthesizes RNA by adding ribonucleotides complementary to the DNA template strand. The reaction can be summarized as:
Reaction:
RNA polymerase reads the DNA template as the duplex unwinds, incorporating complementary nucleotides into the growing RNA chain.
Elongation continues until a termination signal is reached.

Initiation of Transcription
Transcription initiation involves the binding of RNA polymerase to a specific DNA sequence called the promoter. The sigma (σ) subunit of RNA polymerase recognizes and binds to the promoter, guiding the enzyme to the correct initiation site.
Only one DNA strand (the template or antisense strand) is copied.
Synthesis occurs in the 5' to 3' direction relative to the new RNA strand.
The coding (sense) strand has the same sequence as the RNA (except T is replaced by U).

Promoters and Consensus Sequences
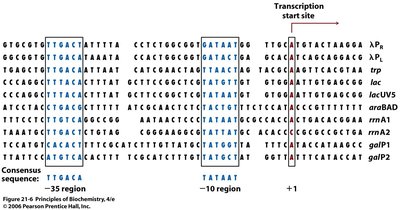
Promoters are DNA sequences upstream of the gene that signal the start of transcription. They contain conserved consensus sequences recognized by RNA polymerase and associated factors.
-10 region (TATA box): TATAAT sequence, located about 10 bases upstream of the transcription start site.
-35 region: TTGACA sequence, located about 35 bases upstream.
Strong promoters: Closely match consensus sequences, leading to higher transcription rates.
Weak promoters: Differ from consensus, resulting in lower transcription rates.
Elongation
During elongation, RNA polymerase moves along the DNA template, synthesizing the RNA strand by adding nucleotides to the 3' end. The DNA double helix unwinds ahead of the polymerase and rewinds behind it.
Termination of Transcription
Termination signals the end of transcription. There are two main mechanisms in prokaryotes:
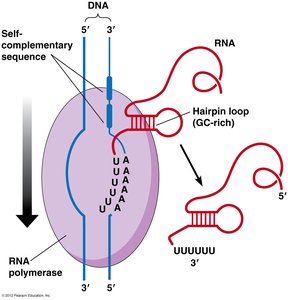
Factor-independent (intrinsic): Involves palindromic, GC-rich sequences that form hairpin loops in the RNA, causing the polymerase to pause and dissociate.
Factor-dependent (Rho-dependent): The Rho protein (a helicase) binds to the nascent RNA and moves toward the polymerase, using ATP to unwind the RNA-DNA hybrid and release the transcript.
Post-Transcriptional Modifications of Eukaryotic mRNA
In eukaryotes, primary mRNA transcripts (pre-mRNA) undergo several modifications before becoming mature mRNA:
5' Capping: Addition of a 7-methylguanosine cap to the 5' end via a 5'-5' triphosphate linkage. This cap protects mRNA from degradation and is essential for ribosome binding during translation initiation.
Polyadenylation (Poly(A) tail): Addition of a stretch of up to 250 adenine nucleotides to the 3' end, enhancing mRNA stability and export from the nucleus.
Splicing: Removal of non-coding sequences (introns) and joining of coding sequences (exons) to produce a continuous coding region.
Regulation of Gene Expression
Gene expression is tightly regulated at multiple levels, including transcriptional and post-transcriptional stages. Regulation ensures that genes are expressed at the right time, place, and amount.
Gene Category | Description |
|---|---|
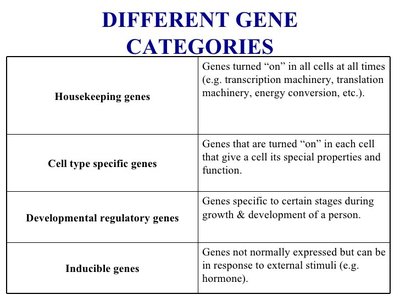
Housekeeping genes | Expressed in all cells at all times; essential for basic cellular functions. |
Cell type specific genes | Expressed only in certain cell types, conferring specialized functions. |
Developmental regulatory genes | Active during specific stages of growth and development. |
Inducible genes | Expressed in response to external stimuli (e.g., hormones). |
Summary of Key Concepts
Genes are DNA segments transcribed into RNA, which may encode proteins or function directly as tRNA or rRNA.
Transcription involves initiation (promoter recognition), elongation (RNA synthesis), and termination (release of RNA transcript).
Post-transcriptional modifications in eukaryotes include 5' capping, polyadenylation, and splicing.
Gene expression is regulated at multiple levels to ensure proper cellular function.
