DNase I cuts DNA that is not protected by bound proteins but is unable to cut DNA that is complexed with proteins. Human DNA is isolated, stripped of its nonhistone proteins, and mixed with DNase I. Samples are removed after 30 minutes, 1 hour, and 4 hours and run separately in gel electrophoresis. The resulting gel is stained to make all DNA fragments in it visible, and the results are shown in the figure. DNA fragment sizes in base pairs (bp) are estimated by the scale to the left of the gel. Examine the gel results and speculate why longer DNase I treatment produces different results.
 Sanders 3rd Edition
Sanders 3rd Edition Ch. 10 - Eukaryotic Chromosome Abnormalities and Molecular Organization
Ch. 10 - Eukaryotic Chromosome Abnormalities and Molecular Organization Problem 27b
Problem 27bGenomic DNA from the nematode worm Caenorhabditis elegans is organized by nucleosomes in the manner typical of eukaryotic genomes, with 145 bp encircling each nucleosome and approximately 55 bp in linker DNA. When C. elegans chromatin is carefully isolated, stripped of nonhistone proteins, and placed in an appropriate buffer, the chromatin decondenses to the 10-nm fiber structure. Suppose researchers mix a sample of 10-nm–fiber chromatin with a large amount of the enzyme DNase I that randomly cleaves DNA in regions not protected by bound protein. Next, they remove the nucleosomes, separate the DNA fragments by gel electrophoresis, and stain all the DNA fragments in the gel.
Explain the origin of DNA fragments seen in the gel.
 Verified step by step guidance
Verified step by step guidance
Verified video answer for a similar problem:
Key Concepts
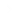
Nucleosome Structure
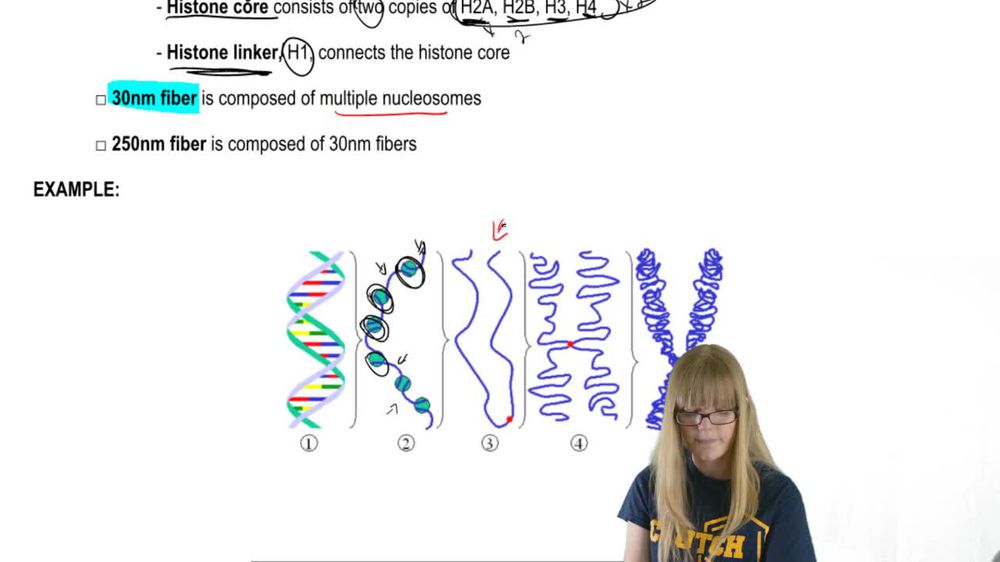
DNase I Activity
Gel Electrophoresis
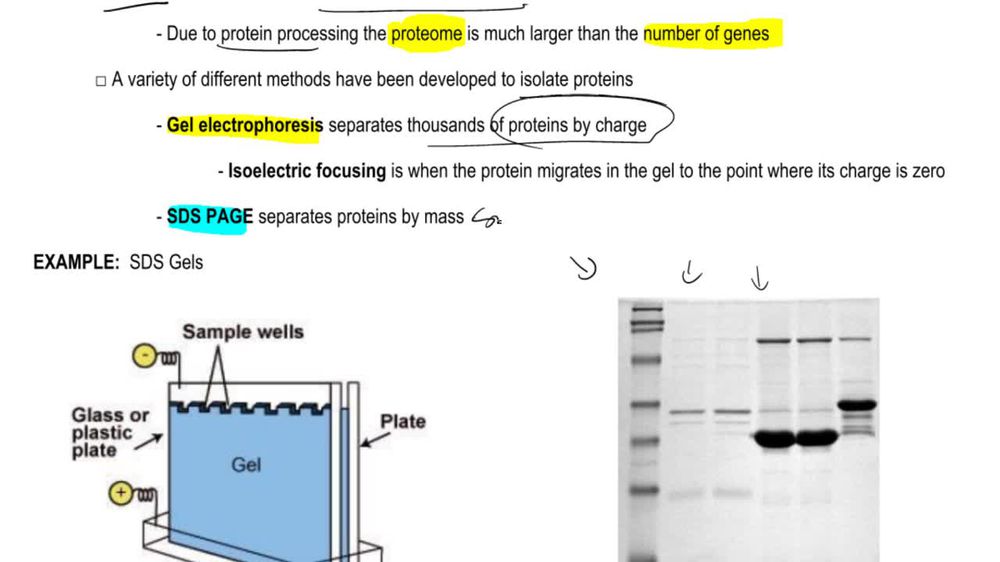

DNase I cuts DNA that is not protected by bound proteins but is unable to cut DNA that is complexed with proteins. Human DNA is isolated, stripped of its nonhistone proteins, and mixed with DNase I. Samples are removed after 30 minutes, 1 hour, and 4 hours and run separately in gel electrophoresis. The resulting gel is stained to make all DNA fragments in it visible, and the results are shown in the figure. DNA fragment sizes in base pairs (bp) are estimated by the scale to the left of the gel. Draw a conclusion about the organization of chromatin in the human genome from this gel.
Genomic DNA from the nematode worm Caenorhabditis elegans is organized by nucleosomes in the manner typical of eukaryotic genomes, with 145 bp encircling each nucleosome and approximately 55 bp in linker DNA. When C. elegans chromatin is carefully isolated, stripped of nonhistone proteins, and placed in an appropriate buffer, the chromatin decondenses to the 10-nm fiber structure. Suppose researchers mix a sample of 10-nm–fiber chromatin with a large amount of the enzyme DNase I that randomly cleaves DNA in regions not protected by bound protein. Next, they remove the nucleosomes, separate the DNA fragments by gel electrophoresis, and stain all the DNA fragments in the gel.
Approximately what range of DNA fragment sizes do you expect to see in the stained electrophoresis gel? How many bands will be visible on the gel?
Genomic DNA from the nematode worm Caenorhabditis elegans is organized by nucleosomes in the manner typical of eukaryotic genomes, with 145 bp encircling each nucleosome and approximately 55 bp in linker DNA. When C. elegans chromatin is carefully isolated, stripped of nonhistone proteins, and placed in an appropriate buffer, the chromatin decondenses to the 10-nm fiber structure. Suppose researchers mix a sample of 10-nm–fiber chromatin with a large amount of the enzyme DNase I that randomly cleaves DNA in regions not protected by bound protein. Next, they remove the nucleosomes, separate the DNA fragments by gel electrophoresis, and stain all the DNA fragments in the gel.
How do the expected results support the 10-nm–fiber model of chromatin?
A small population of deer living on an isolated island is separated for many generations from a mainland deer population. The populations retain the same number of chromosomes but hybrids are infertile. One chromosome (shown here) has a different banding pattern in the island population than in the mainland population.
Describe how the banding pattern of the island population chromosome most likely evolved from the mainland chromosome. What term or terms describe the difference between these chromosomes?
A small population of deer living on an isolated island is separated for many generations from a mainland deer population. The populations retain the same number of chromosomes but hybrids are infertile. One chromosome (shown here) has a different banding pattern in the island population than in the mainland population.
Draw the synapsis of these homologs during prophase I in hybrids produced from the cross of mainland with island deer.