 Back
BackAmino Acids, Peptide Bonds, and Protein Purification: Study Notes for Biochemistry
Study Guide - Smart Notes

Amino Acids and Peptide Bonds
Peptide Bond Resonance
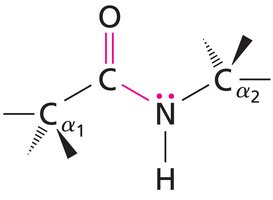
The peptide bond is a fundamental linkage between amino acids in proteins, formed by a condensation reaction between the carboxyl group of one amino acid and the amino group of another. Peptide bonds exhibit partial double bond character due to resonance, which restricts rotation and maintains planarity.
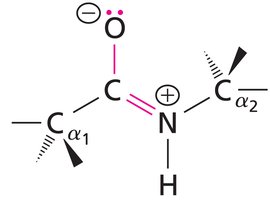
Resonance Structure: The peptide bond alternates between a single bond and a double bond configuration, resulting in a planar structure.
Implications: This planarity is crucial for protein secondary structure formation, such as alpha helices and beta sheets.
Example: The resonance forms of the peptide bond are shown below.
Ionizable Groups and pKa Values
Amino acids contain ionizable groups whose charge state depends on the pH of the environment. The pKa values of these groups determine their protonation state, which is essential for understanding peptide charge and protein function.
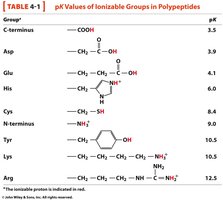
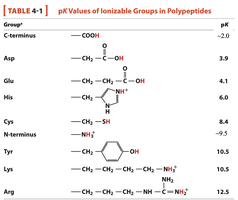
Key Ionizable Groups: C-terminus, N-terminus, side chains of Asp, Glu, His, Cys, Tyr, Lys, Arg.
pKa Values: Each group has a characteristic pKa, which can be used to predict its charge at a given pH.
Application: Calculating net charge and formal charge of peptides at different pH values.
Group | Structure | pKa |
|---|---|---|
C-terminus | COOH | ~2.0-3.5 |
Asp | CH2-COOH | 3.9 |
Glu | CH2-CH2-COOH | 4.1 |
His | Imidazole | 6.0 |
Cys | CH2-SH | 8.4 |
N-terminus | NH3+ | ~9.5-10.5 |
Tyr | Phenol | 10.5 |
Lys | CH2-CH2-CH2-CH2-NH3+ | 10.5 |
Arg | CH2-CH2-CH2-NH-C(NH2)2+ | 12.5 |
Peptide Charge States and Calculations
Charge State Determination
The charge state of a peptide depends on the pH relative to the pKa of its ionizable groups. At physiological pH (~7), acidic groups are deprotonated (negative), basic groups are protonated (positive), and the microenvironment can affect pKa values.
Rule: If pH < pKa, the group is protonated; if pH > pKa, the group is deprotonated.
Calculation: Formal charge and net charge can be determined by summing the charges of all ionizable groups.
Example: For a peptide, calculate the net charge at pH 7 using the pKa values and the formula:
Protein Purification and Separation Techniques
Overview of Protein Purification
Protein purification is essential for studying protein structure and function. It involves isolating a target protein from a complex mixture using various separation techniques based on physical and chemical properties.
Steps: Molecular cloning and overexpression, cell lysis, selective separation, and purification.
Properties Used: Solubility, ionic charge, hydrophobicity, size, and binding specificity.
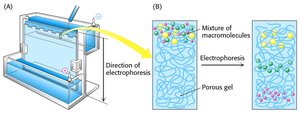
Polyacrylamide Gel Electrophoresis (PAGE)
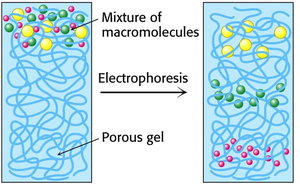
PAGE is a technique used to separate proteins based on their size, charge, and shape. The gel is made from polymerized acrylamide, and the density of the gel determines the size range for protein separation.
Native PAGE: Maintains protein complexes and associations; used to study protein-protein or protein-DNA interactions.
Denaturing PAGE (SDS-PAGE): Uses SDS to denature proteins and impart a uniform negative charge, so separation depends only on size.
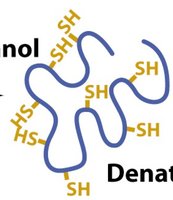
Reducing Agents: DTT, BME, and TCEP are used to break disulfide bonds, further denaturing proteins.
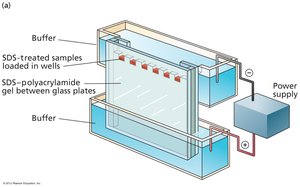
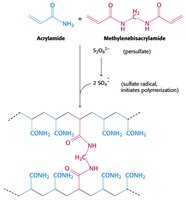
SDS-PAGE Sample Preparation and Visualization
SDS-PAGE requires denaturation of proteins using SDS and heat, followed by reduction of disulfide bonds. Visualization is achieved using Coomassie Brilliant Blue dye or Western blotting for specific protein detection.
SDS: Disrupts tertiary structure and imparts uniform negative charge.
Reducing Agents: Break disulfide bonds for complete denaturation.
Visualization: Coomassie dye stains all proteins; Western blotting detects specific proteins using antibodies.
Determining Molecular Weight by SDS-PAGE
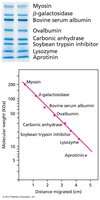
The molecular weight of proteins can be determined by comparing the migration distance of unknown proteins to standards and plotting against the log of molecular weight.
Standard Curve: Plot distance migrated vs. log(MW) for standards.
Unknowns: Measure migration distance, locate on curve, and determine MW.

Chromatography Techniques for Protein Purification
Column Chromatography Principles
Column chromatography separates proteins based on charge, size, or binding affinity. The stationary phase is a solid matrix, and the mobile phase is a buffer or solvent.
Types: Ion exchange, size exclusion, affinity, and reverse-phase chromatography.
Elution: Proteins are washed off the column by changing buffer conditions.
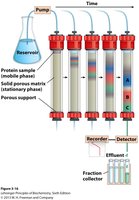
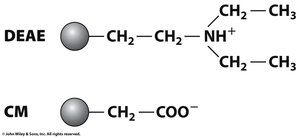
Ion Exchange Chromatography
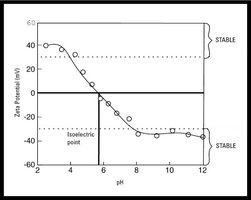
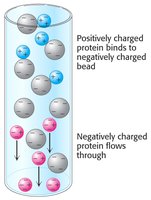
Ion exchange chromatography separates proteins based on their net charge at a given pH. The isoelectric point (pI) is the pH at which a protein has no net charge.
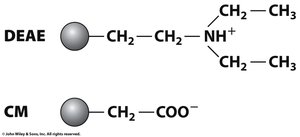
Cation Exchange: Positively charged proteins bind to negatively charged matrices.
Anion Exchange: Negatively charged proteins bind to positively charged matrices.
Elution: Increasing salt concentration elutes proteins from the column.
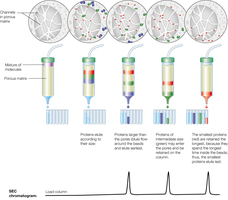
Size Exclusion Chromatography (SEC)
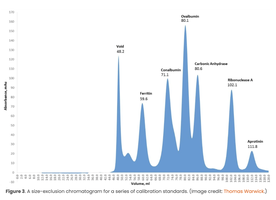
SEC separates proteins based on size and shape. Beads with defined pore sizes allow smaller proteins to enter and elute later, while larger proteins elute earlier.
Rule of Thumb: Useful for separating proteins with at least a 2x size ratio.
Application: Determining molecular weight and studying protein complexes.

Affinity Chromatography
Affinity chromatography separates proteins based on specific binding interactions with a matrix decorated with a chemical group. Genetically encoded affinity tags are commonly used for purification.
Tags: Polyhistidine (His-tag), MBP, GST.
Elution: Addition of a competitor (e.g., imidazole for His-tag) releases the target protein.
Application: Highly specific purification of recombinant proteins.
Summary of Key Concepts
Memorize amino acid names, codes, structures, and properties.
Understand peptide bond formation and resonance.
Calculate charge states and pI for peptides and proteins.
Apply SDS-PAGE and native PAGE for protein analysis.
Utilize ion exchange, size exclusion, and affinity chromatography for protein purification.
Additional info: These notes expand on brief points from the original materials, providing academic context and examples for biochemistry students.
