 Back
BackThe Genetic Code and Translation: Structure, Function, and Mechanisms
Study Guide - Smart Notes

Topic 14: The Genetic Code and Translation
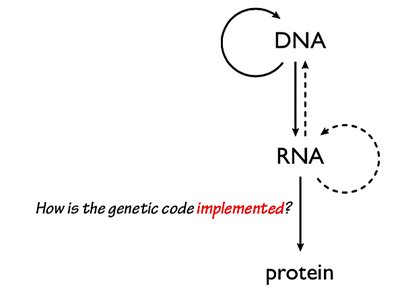
The Central Dogma of Molecular Biology
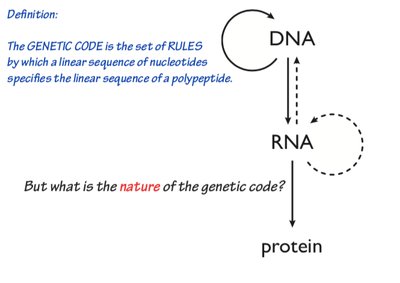
The central dogma describes the flow of genetic information within a biological system: DNA is transcribed into RNA, which is then translated into protein. This process underlies all cellular function and heredity.
DNA Replication: The process by which DNA makes a copy of itself.
Transcription: The synthesis of RNA from a DNA template.
Translation: The synthesis of proteins from an mRNA template.
The Genetic Code: Structure and Properties
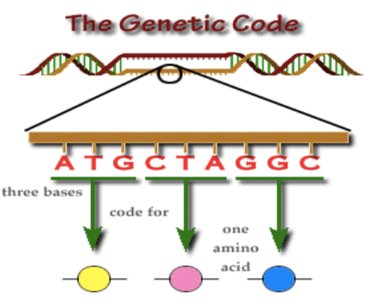
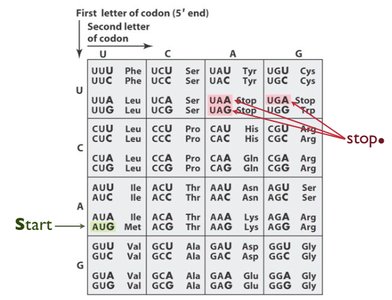
The genetic code is a set of rules by which the sequence of nucleotides in mRNA is translated into the sequence of amino acids in proteins. It is composed of codons, which are groups of three nucleotides that specify particular amino acids.
Codon: A sequence of three nucleotides in mRNA that codes for a specific amino acid or a stop signal.
Triplet Code: Each codon consists of three bases, allowing for 64 possible combinations (43 = 64).
Reading Frame: The way nucleotides are grouped into codons; only one reading frame is correct for a given mRNA.
Non-overlapping: Each nucleotide is part of only one codon.
Universal: The genetic code is nearly universal among all organisms, with only minor exceptions (e.g., mitochondria).
Degenerate: Most amino acids are encoded by more than one codon.
Start Codon: AUG (methionine) signals the start of translation.
Stop Codons: UAA, UAG, and UGA signal the end of translation.
Discovery of the Genetic Code
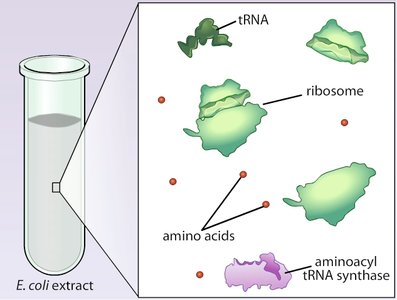
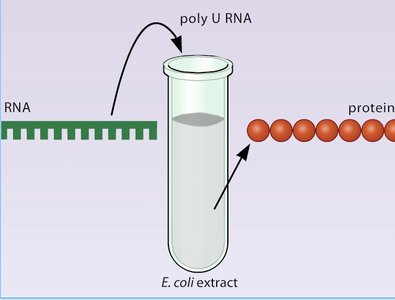
The genetic code was deciphered through experiments using cell-free extracts and synthetic RNAs. Marshall Nirenberg and colleagues demonstrated that synthetic RNA sequences could direct the synthesis of specific polypeptides, revealing the correspondence between codons and amino acids.
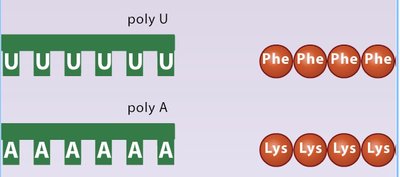
Poly-U Experiment: Polyuridylic acid (poly-U) directed the synthesis of polyphenylalanine, showing that UUU codes for phenylalanine.
Further Experiments: Other homopolymers (poly-A, poly-C) were used to identify additional codon assignments.


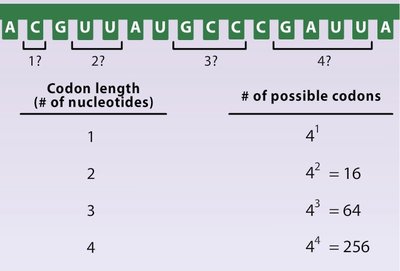
Why a Three-Letter Code?
There are four nucleotide bases (A, U, G, C) and 20 amino acids. A single base code would only allow for 4 amino acids, and a doublet code for 16. A triplet code allows for 64 combinations, more than enough to encode all 20 amino acids.
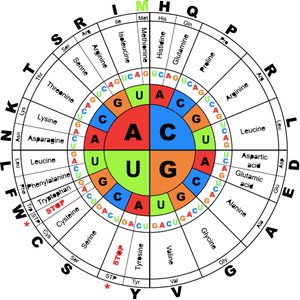
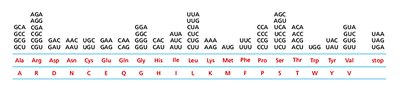
Codon Assignments and Degeneracy
Most amino acids are specified by more than one codon, a feature known as degeneracy. Codons for the same amino acid often share the same first two bases and differ at the third position.
Start Codon: AUG (methionine in eukaryotes, formylmethionine in prokaryotes).
Stop Codons: UAA (ochre), UAG (amber), UGA (opal).
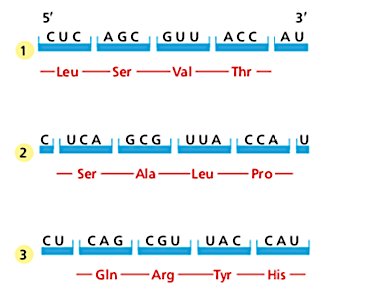
Reading Frames and the Importance of AUG
The reading frame is established by the first AUG codon. Shifting the reading frame by one or two nucleotides changes the resulting amino acid sequence entirely, emphasizing the importance of correct initiation.
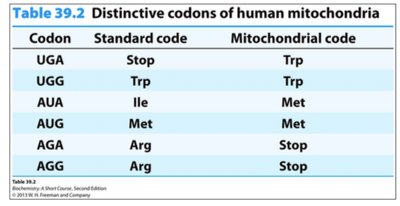
Universality and Variations of the Genetic Code
The genetic code is nearly universal, but some variations exist, such as in human mitochondria, where certain codons are reassigned.
Codon | Standard code | Mitochondrial code |
|---|---|---|
UGA | Stop | Trp |
UGG | Trp | Trp |
AUA | Ile | Met |
AUG | Met | Met |
AGA | Arg | Stop |
AGG | Arg | Stop |
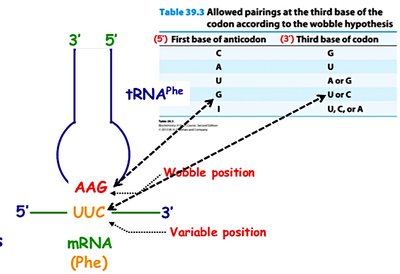
The Wobble Hypothesis
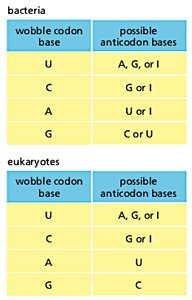
The wobble hypothesis explains how a single tRNA can recognize more than one codon. The third base of the codon (5' end of the anticodon) allows for non-standard base pairing, increasing the efficiency of translation.
Wobble Position: The 5' base of the tRNA anticodon pairs flexibly with the 3' base of the mRNA codon.
Inosine (I): A modified base in tRNA that can pair with U, C, or A.
Wobble codon base | Possible anticodon bases (bacteria) | Possible anticodon bases (eukaryotes) |
|---|---|---|
U | A, G, or I | A, G, or I |
C | G or I | G or I |
A | U or I | U |
G | C or U | C |
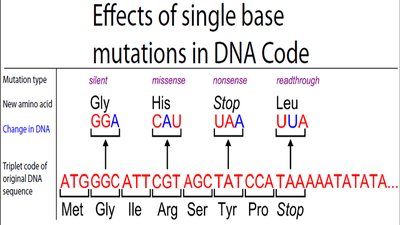
Mutations and the Genetic Code
Mutations are changes in the nucleotide sequence that can alter the amino acid sequence of proteins. Types include silent, missense, nonsense, and readthrough mutations.
Silent Mutation: No change in amino acid sequence.
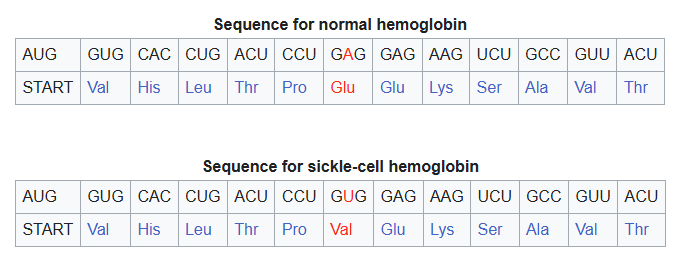
Missense Mutation: One amino acid is replaced by another.
Nonsense Mutation: A codon is changed to a stop codon, truncating the protein.
Readthrough Mutation: A stop codon is changed to code for an amino acid, extending the protein.

Translation: Mechanism and Components
Overview of Translation
Translation is the process by which the information in mRNA is used to synthesize proteins. It occurs in the cytoplasm and involves ribosomes, tRNAs, and various accessory factors.
mRNA: Provides the template for protein synthesis.
tRNA: Adaptor molecules that bring amino acids to the ribosome.
Ribosome: The molecular machine that catalyzes peptide bond formation.
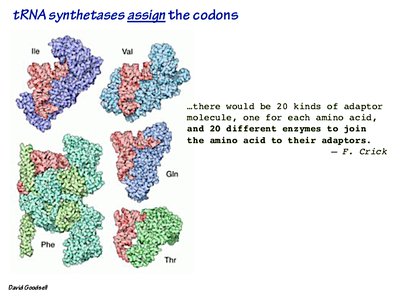
Aminoacyl-tRNA synthetases: Enzymes that attach amino acids to their corresponding tRNAs.
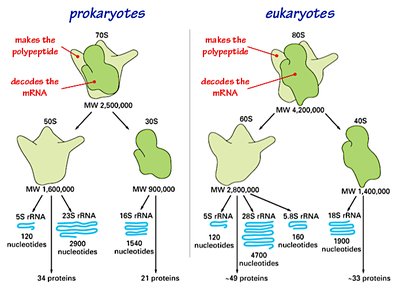
Ribosome Structure and Function
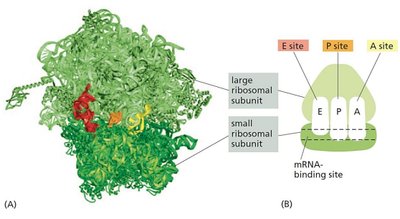
Ribosomes are composed of two subunits (large and small), each containing rRNA and proteins. They have three binding sites for tRNA: A (aminoacyl), P (peptidyl), and E (exit).
Small Subunit: Decodes the mRNA.
Large Subunit: Catalyzes peptide bond formation.
rRNA: Forms the catalytic core of the ribosome.
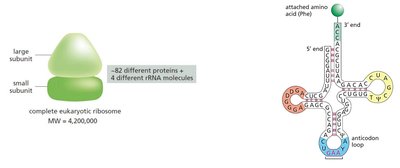
tRNA: Structure and Function
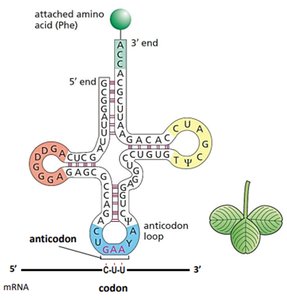
tRNAs are small RNA molecules (~75-95 nucleotides) with a cloverleaf secondary structure. Each tRNA has an anticodon that pairs with a codon in mRNA and a 3' end that attaches to a specific amino acid.
Anticodon Loop: Contains the three-nucleotide sequence complementary to the mRNA codon.
3' CCA End: Site of amino acid attachment.
Aminoacyl-tRNA Synthetase: Enzyme that charges tRNA with its corresponding amino acid.
Stages of Translation
Translation occurs in three main stages: initiation, elongation, and termination.
Initiation: Assembly of the ribosome on the mRNA at the start codon.
Elongation: Sequential addition of amino acids to the growing polypeptide chain.
Termination: Release of the completed polypeptide when a stop codon is encountered.
Initiation: Prokaryotes vs Eukaryotes
Initiation differs between prokaryotes and eukaryotes, particularly in how the ribosome recognizes the start site.
Prokaryotes: The small ribosomal subunit binds directly to the Shine-Dalgarno sequence upstream of the start codon.
Eukaryotes: The small subunit binds to the 5' cap and scans for the first AUG codon.
Elongation
During elongation, aminoacyl-tRNAs enter the A site, peptide bonds are formed, and the ribosome translocates along the mRNA.
Peptidyl Transferase: Catalyzes peptide bond formation in the large subunit.
Translocation: Ribosome moves one codon downstream, shifting tRNAs between sites.
Termination
Termination occurs when a stop codon enters the A site. Release factors bind, prompting the release of the polypeptide and dissociation of the ribosome.
Stop Codons: UAA, UAG, UGA.
Release Factors: Proteins that recognize stop codons and trigger termination.
Polysomes
Multiple ribosomes can translate a single mRNA simultaneously, forming a structure called a polysome. This increases the efficiency of protein synthesis.
Protein Synthesis Inhibitors
Certain antibiotics and toxins can inhibit protein synthesis by targeting the ribosome or translation factors, which is important in both medicine and research.
Summary Table: Key Features of the Genetic Code
Feature | Description |
|---|---|
Triplet Code | Three nucleotides per codon |
Degeneracy | Most amino acids have multiple codons |
Start Codon | AUG (methionine) |
Stop Codons | UAA, UAG, UGA |
Universality | Nearly universal, with minor exceptions |
Non-overlapping | Each nucleotide is part of only one codon |
Wobble | Flexibility at the third codon position |
Additional info: The notes above integrate foundational biochemistry concepts with experimental evidence and molecular mechanisms, providing a comprehensive overview suitable for college-level study.
